Escolar Documentos
Profissional Documentos
Cultura Documentos
Enzyme Performance Factors
Enviado por
Mikkel EliasenDescrição original:
Título original
Direitos autorais
Formatos disponíveis
Compartilhar este documento
Compartilhar ou incorporar documento
Você considera este documento útil?
Este conteúdo é inapropriado?
Denunciar este documentoDireitos autorais:
Formatos disponíveis
Enzyme Performance Factors
Enviado por
Mikkel EliasenDireitos autorais:
Formatos disponíveis
Applied
Statistics and Statistical Software, 02441
Effect of hardness and detergent on enzymatic catalysis
Table of Contents
Technical University of Denmark Dept of Mathematics and Computer
Summary .............................................................................................................................................. 1 Introduction .......................................................................................................................................... 1 Data ...................................................................................................................................................... 2 Methods ................................................................................................................................................ 3 Results .................................................................................................................................................. 4 Discussion ............................................................................................................................................ 6 Conclusion ........................................................................................................................................... 7 Appendix I............................................................................................................................................ 8 Appendix II .......................................................................................................................................... 9
Technical University of Denmark Dept of Mathematics and Computer
1 2 3 4 5 6 7 8 9 10 11
Summary
Different
enzymes
improve
textile
washing,
and
it
is
of
interest
to
know
performance
under
various
conditions.
The
performance
of
five
enzymes
was
measured
as
the
amount
of
protein
removed
from
a
surface
when
exposed
to
the
factors
hardness
and
detergent.
We
construct
a
model
using
variance
analysis
that
accounts
for
90%
of
the
variance
in
the
data.
We
show
that
hardness
of
water
has
no
significant
influence
on
enzyme
performance,
and
that
addition
of
detergent
yields
the
highest
protein
removal.
Adding
more
protein
to
samples
also
has
an
effect
on
protein
removal,
but
the
effect
is
saturating.
Using
the
15
nM
enzyme
concentration
and
addition
of
detergent
shows
that
protein
A
removes
proteins
of
surfaces
significantly
better
than
the
other
proteins
included.
We
conclude
that
addition
of
detergent,
concentration
of
enzymes
and
enzyme
type
is
of
significant
importance
in
stain
removal
processes.
12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28
Introduction
Efficient
protein
removal
is
of
vital
importance
in
the
textile
cleaning
industry.
Enzymatic
activity
enhances
this
process
by
means
of
catalyzation.
It
is
known
that
factors,
such
as
detergent
and
water
hardness
affect
this
catalyzation
rate,
and
it
is
therefore
of
interest
to
quantify
these
effects.
These
factors
are
some
of
the
conditions
that
appear
in
normal
laundry
wash
processes.
To
optimize
the
effect
of
removing
certain
stains
on
textile
surfaces
the
effect
is
analyzed
by
means
of
a
laboratory
experiment.
The
experiment
measures
how
much
protein
is
removed
from
a
surface
with
Surface
Plasmon
Resonance
technology
(SPR),
which
is
a
biosensor
that
measures
a
resonance
signal
on
a
gold
surface,
and
translates
it
into
a
protein
removal
response.
The
experiment
is
conducted
using
5
different
proteins
under
different
conditions.
The
conditions
are
addition
of
detergent,
hard
and
soft
water,
and
a
range
of
enzymatic
concentrations.
A
replication
of
each
experiment
yields
a
grand
total
of
160
different
samples.
We
wish
to
use
statistical
methods
to
examine
differences
and
trends
in
the
samples.
The
aim
of
the
analysis
is
to
describe
and
compare
the
performance
of
the
enzymes
and
how
different
conditions
affect
enzyme
performance.
This
analysis
includes
factorial
interactions,
determining
whether
the
day
to
day
measurements
are
reliable
and
testing
robustness
of
the
experimental
set-up.
1|Page
Technical University of Denmark Dept of Mathematics and Computer
29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52
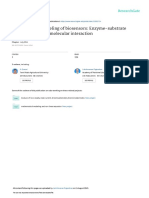
Figure 1: A) Boxplot of protein response with detergent+ and detergent0. Detergent plus alone seems to have a positive effect on the response. B) The response on hardness of water. There does not seem to be a correlation between hardness and response.
Data
The
dataset
consists
of
160
samples,
with
80
different
experimental
combinations.
The
enzymes
were
labeled
A,
B,
C,
D
and
E
and
analyzed
one
at
a
time,
with
the
experiment
running
for
2
consecutive
days.
No
enzymes
were
run
the
same
day,
so
each
specific
enzyme
is
labeled
with
a
single
date.
After
the
experiment
has
run,
the
result
is
sampled
randomly
into
a
variable
called
cycles.
The
output
is
response
which
is
given
as
the
amount
of
protein
removed
in
RU
(10-6
g
m-2).
The
factors
included
in
the
data
are
hardness
of
water
and
addition
of
detergent.
Both
these
variables
are
binary
and
denoted
De+,
De0,
Ca+
and
Ca0,
for
detergent
and
calcium
respectively.
Figure
1
shows
the
log10(Response)
together
with
detergent
and
hardness.
The
enzyme
concentrations
were
tested
under
4
different
levels:
0
nM,
2.5
nM,
7.5
nM
and
15
nM.
Increasing
the
concentration
of
enzymes
in
the
samples
also
increases
the
response.
There
is
some
indication
in
the
data
that
the
enzyme
catalyzation
gets
saturated
at
the
high
concentrations
(Figure 2).
This
may
potentially
yield
two
problems:
fitting
the
data
as
a
linear
model
gives
more
imprecise
results
at
high
concentrations,
and
Figure 2: Response vs concentration of the 5 different enzymes. Black = 0 nM, Red = 2.5 nM, Green = 7.5 nM and Blue = 15 nM. Increasing concentration increases the response, but there is some saturation towards the high concentrations.
extrapolation beyond existing data points is very uncertain. A summary of all data included in our analysis can be seen in Table 1. 2|Page
Technical University of Denmark Dept of Mathematics and Computer
53 54 55
Table 1: Table of variables and response variable in the model. With Detergent (Det+) or without (Det0). Hardness: with (Ca+) or without (Ca0)
Variable Name Variables Run Date Cycle Enzyme Type Enzyme Concentration Detergent Hardness Output Response
Type Categorical Categorical Categorical Continuous Categorical Categorical Continuous
Mean
Categories 3/12/2008 5/12/2008 1, 2, 3 34 A, B, C, D, E
Variance 33.0 154254.4
Levels 10 34 5
6.3
Det+, Det0 Ca+, Ca0
2 2
434.3
56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74
Methods
Our first approach was to assume the level of response is given as a function of the various factors and enzymes. Initially we assumed that there was no bias in the response measurements for neither cycle nor days. Concentration was used as a continuous variable, as this allowed us to interpolate between levels, if this is needed in the future. An ANCOVA analysis of data was conducted using two-way interactions. The equation for ANCOVA is
!" = ! + ! ! + !"
(1)
Where Y is the Response variable, i is the intercept of the model with factor i. is the slope in the model and Xj are the observations in the data. is noise. When subsetting data and only test for one factorial variable we use a regular ANOVA !" = ! + !" (2)
The assumptions for these models are that residuals are independent, have constant variance and are normal distributed. To verify this we inspected the fit with residuals, a QQ-normality plot, and Cooks distance for the observations (Appendix 1). These figures revealed some inconsistencies in variance. In order to deal with this we log transformed the response variable before making the model. This gave acceptable distributions, with the exception of a single outlier: Enzyme B, 3|Page
Technical University of Denmark Dept of Mathematics and Computer
75 76 77 78
concentration = 0, Det0 and Ca0 looks biased in the response, and was thus removed from the data set. All statistical analysis was performed using R 2.14.1, scripts and analysis are documented in Appendix II.
79 80 81 82 83 84 85 86 87 88 89 90
Results
We tested a multi-way ANCOVA model with log-response as a function of the continuous variable concentration, and the 3 categorical variables Enzyme, Detergent, Hardness and all their two way interactions. The model showed that Hardness had no significant effect (p > 0.29) on response. After removing Hardness as a factor the model showed that the two-way interaction between Enzyme type and Enzyme concentration had no effect either (p > 0.30). This concludes that detergent, enzyme type and concentration significantly influence protein removal. The sum of squares value shows that the most important factor to remove protein is detergent. The final model explains 89% of the variation in the data. P-values and a summary of the model are shown in table II.
Table 2: Results from the ANCOVA model including Enzyme type, Enzyme Concentration and Detergent. The interactions are denoted with a :. Significance levels: *** p < 0.001, ** p < 0.01, * p < 0.5.
Enzyme Type Enzyme Concentration Detergent Enzyme:Detergent Enzyme Concentration:Detergent
Degrees of Sum of P-value freedom squares 4 1 1 4 1 3,932 15,611 31,327 0,711 2,768 1,33E-14*** 2,33E-27*** 1,31E-44*** 3,54E-03** 3,61E-13***
91 92 93 94 95 96 97 Secondly we wanted to determine whether or not the interactions where the same for all enzymes included in the model. As we already concluded that detergent significantly influenced data, we used a more graphical approach. Figure 3 shows that some of the enzymes react slightly different to addition of detergent. As the difference between the curves are not the same with and without detergent implies that not all the proteins interact the same way with the factor. Enzyme B is apparently a little less efficient relative to the others when detergent is added.
4|Page
Technical University of Denmark Dept of Mathematics and Computer
Figure 3: log10 response vs enzyme concentration for the 5 different enzymes. A) with detergent added to the sample, B) without detergent added. Structural differences between the two plots suggest stronger interactions between some enzymes with detergent than others.
98 99 100 101 102 103 104 105 106 107 108
The next step was to investigate if one enzyme was significantly removing proteins better than the others. To do so we split up the data set into a smaller group, with the optimal conditions for protein removal; with detergent added and 15nM enzyme added. With an ANOVA-test we were able to determine that enzyme A was the best enzyme with a mean protein removal of 1527.3 RU, with a possible minimum of 1391.7 and a maximum of 1676.2 on a 95% confidence interval. In comparison the worst enzyme was D with a mean protein removal of 663.5 RU. The summary of all the enzymes can be seen in Table 3, which also states the p-values compared to enzyme A.
Table 3 The 5 enzymes compared to enzyme A with 15nM added. ANOVA shows that all the other enzymes are significantly different from A. Given is also the mean and the 95% confidence interval. Significance levels: *** p < 0.001, ** p < 0.01, * p < 0.5.
Enzyme A Enzyme B Enzyme C Enzyme D Enzyme E
Estimate 1527 980 1023 663 1101
2.5% 1392 893 932 605 1003
97.5% 1676 1076 1122 728 1208
P-value 3,15E-06 *** 1,00E-05 *** 8,39E-10 *** 8,87E-05 ***
109 110 111 112 113 114 5|Page Analysis of experimental bias To investigate if there was any systematic error in the data sampling, we also tested the response compared to the random variable, cycle. Furthermore we tested day to day variation by testing the samples which has 0 nM concentration of enzymes against one another.
Technical University of Denmark Dept of Mathematics and Computer
115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130
To test if cycle sampling influenced the deterioration of proteins we made an ANOVA with cycle as explanatory variable. As all cycles are sampled randomly, there should be no significant difference between the cycles. This turns out to be true as the difference between cycles is insignificant (p > 0.47). The day-to-day runs were explored by an ANCOVA analysis, using hardness, Enzyme type and detergent as explanatory variables, and with concentration 0. The analysis shows that enzyme types are significantly different (p < 0.0013). As the samples should be equal (no enzyme added), this strongly indicates experimental bias in the setup, or sampling error at the day enzyme B was analyzed. Using concentration as factor instead of continuous variable Enzyme concentration was given at four different levels, which yielded the possibility of using the variable as a factor or as a continuous variable. In the previous section we have used it as a factor, in order to be able to extrapolate the model. Making a model with concentration as a factor, and comparing it to a model with concentration as a continuous variable , and comparing them with an ANOVA shows that using concentration as a factor gives a significantly better fit (p < 1.31E-21). The implication and possible solutions of this are evaluated in the discussion.
131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147
Discussion
The few data points for each group give large statistical uncertainty in the validation of the data, and therefore the credibility of the results. More data would have given us more samples to do the statistically analysis and check for errors within the different groups. Notably the sample or experimental bias within day B yields high uncertainty. With the data set available it would be reasonable to adjust data to an expected mean with 0 enzyme added. The addition of the reference samples would also be helpful in this particular problem. In order for our analysis to be valid in terms of hardness and detergent in the water, we assume that the concentrations of these two factors are the same in all observations. Concentration was used as a continuous variable instead of a factor, in spite of the fact that using it as a factor gave a significantly better model. This was done in order to be able to interpolate between experiments with different concentration. In order to make a better model it should be considered to make a model more appropriate than a linear fit for enzyme kinetics, which apparently looks more like a Michaelis-Menten saturation model. If the main purpose of the experiment is to determine the catalytic capabilities of enzymes, the addition of less detergent would be appropriate, as it turns out to be the main factor explaining protein degradation from surfaces. 6|Page
Technical University of Denmark Dept of Mathematics and Computer
148 149 150
Future work with the data would include a more thorough analysis of interactions with the individual enzymes. All five types of enzyme have a significant reaction with detergent, as well as with concentration, but to what degree is important knowledge.
151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178
Conclusion
We conclude that hardness does not have an effect on protein degradation in our experiment. Oppositely detergent, enzyme concentration and enzyme type has a highly significant positive effect. Their interactions are also significant, with the interaction between concentration and enzyme type omitted. We also conclude that enzyme A has the highest catalytic capabilities under optimal conditions, and that enzyme D is the worst enzyme for removing proteins.
7|Page
179 180
Appendix
I
Figure
1
shows
the
residuals
of
the
log
transformed
analysis.
Technical University of Denmark Dept of Mathematics and Computer
181 182 183 184 185 186 187 188
8|Page
189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228
Appendix
II
R-code used to calculate statistics in the report.
# Script for case 1 detergent
Technical University of Denmark Dept of Mathematics and Computer
################################# Load and view data ########################### setwd("~/My Dropbox/02441 Applied Statistics and Statistical Software/Case_1") # Set working directory graphics.off()
mydata <- read.table('SPR.txt', header = TRUE, colClasses = c('factor', 'factor','numeric','factor', 'numeric','factor','factor'))
#################################### Manipulate data for cleaner analysis ######################### mydata$Cycle <- factor(mydata$Cycle, levels = 1:34) # Put cycles conscutively mydata$lResponse <- log10(mydata[,3]) # Log response for equal variance mydata <- mydata[-14,] # Remove outlier 14 mydata$EnzymeConcF <- factor(mydata$EnzymeConc ,levels = c('15','7.5','2.5','0')) # Make Enzyme a factor for extra model determination ##################################### Make a linear model of all variables ####################### fm <- lm(lResponse ~ (Enzyme+EnzymeConc+DetStock+CaStock)^2 , data = mydata) m <- summary(fm) # See and assign summary anova(fm) # Make ANCOVA to see #### Leave calcium out of the final model, also the interaction betweeen EnzCon and Enz #####
fm1 <- lm(lResponse ~ (Enzyme+EnzymeConc+DetStock)^2 - Enzyme:EnzymeConc, data = mydata) summary(fm1) an <- anova(fm1) an ##### Use concentration as a factor to compare the two models ######## fmfac <- lm(lResponse ~ (Enzyme+EnzymeConcF+DetStock)^2-Enzyme:EnzymeConcF, data = mydata) summary(fmfac) aF <- anova(fmfac) aF anf <- anova(fm1,fmfac, test = 'F') anf #### Using Concentration as a factor describes data significantly better #######################
#### Show graphically that the data have equal variance and are normally distributed ####
9|Page
229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269
Technical University of Denmark Dept of Mathematics and Computer
graphics.off() # Remove existing graphics windows(width = 7) # Make a new window 7 inches wide par(mfrow=c(2,2)) # 4 subplots, in a 2by2 matrix par(mar = c(5,4,2,1)) # Make margins as small as possible plot(fm1, which = 1:4) # plot the model, with simplified cooks distance
################################ See the response as a function of enzyme concentration ##### graphics.off() cols <- 1:5 plot(lResponse ~ EnzymeConc, data = mydata, type = 'n') points(lResponse ~ EnzymeConc, data = mydata,subset = mydata[,4] == 'A', col = cols[1]) #abline(afm[1], afm[2]) points(lResponse ~ EnzymeConc, data = mydata,subset = mydata[,4] == 'B', col = cols[2]) points(lResponse ~ EnzymeConc, data = mydata,subset = mydata[,4] == 'C', col = cols[3]) points(lResponse ~ EnzymeConc, data = mydata,subset = mydata[,4] == 'D', col = cols[4]) points(lResponse ~ EnzymeConc, data = mydata,subset = mydata[,4] == 'E', col = cols[5]) ############################ Looks as a saturating function #################################
########################## Test if the days differ with 0 protein added ##################### fm0 <- lm(lResponse ~ (Enzyme+CaStock+DetStock)^2, data = mydata ,subset = mydata[,5] == '0') summary(fm0) anova(fm0) ############################ There is significantly differences between the days with 0 enzyme ###
########################## See if the enzymes differ under optimal conditions #################### mydatadet <- mydata[mydata[,6] == 'Det+',] # Find all the data with Det+ fmHigh <- lm(lResponse ~ Enzyme -1, data = mydatadet, subset = mydatadet[,5] == '15') # Make a linear model, and get all intercepts (remove -1 for more usable model). Subset 15 nM p <- summary(fmHigh) # Put summary into a variable ints <- confint(fmHigh) # Calculate the 95% confidence anova(fmHigh) # There is difference among enzymes mens <- fmHigh$coefficient # Retrieve means
####################### A is significantly better than all the other enzymes ######################
####################### Do the enzymes interact the same way between effects ######### enzA <- mydata[mydata[,4] == 'A',] fmA <- lm(lResponse ~ (DetStock + EnzymeConc)^2, data = enzA) summary(fmA) anova(fmA)
10 | P a g e
270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310
Technical University of Denmark Dept of Mathematics and Computer
enzB <- mydata[mydata[,4] == 'B',] fmB <- lm(lResponse ~ (DetStock + EnzymeConc)^2, data = enzB) summary(fmB) anova(fmB)
enzC <- mydata[mydata[,4] == 'C',] fmC <- lm(lResponse ~ (DetStock + EnzymeConc)^2, data = enzC) summary(fmC) anova(fmC)
enzD <- mydata[mydata[,4] == 'D',] fmD <- lm(lResponse ~ (DetStock + EnzymeConc)^2, data = enzD) summary(fmD) anova(fmD)
enzE <- mydata[mydata[,4] == 'E',] fmE <- lm(lResponse ~ (DetStock + EnzymeConc)^2, data = enzE) summary(fmE) anova(fmE) ############################ They are all significantly correlated with both effects ########
########################### Plot different figures ##########################################
graphics.off() cols <- 1:4 windows(width = 3.5,height= 2) par(mar=c(2.5,4.5,0.8,0.5)) dat <- tapply(mydata$lResponse,list(mydata$EnzymeConc,mydata$Enzyme),mean) barplot(dat, beside = TRUE, xlab = 'Enzyme' , ylab = 'log10 Response', col = cols, ylim = c(0,3)) abline(0,0) # Bar plot of concentration vs response for all 5 enzymes
#################### Detergent figures ####################################### mydatadet <- mydata[mydata[,6] == 'Det+',] lvl <- levels(mydata$EnzymeConcF) enZ <- levels(mydata$Enzyme) Detline <- matrix(0,4,5) for (i in 1:4)
11 | P a g e
311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351
Technical University of Denmark Dept of Mathematics and Computer
for (j in 1:5){{ sub1 <- mydatadet[mydatadet[,5] ==lvl[i],] Detline[i,j] <- mean(subset(sub1$lResponse,sub1[,4] == enZ[j])) }} rownames(Detline) <- lvl colnames(Detline) <- enZ
mydatadet <- mydata[mydata[,6] == 'Det0',] Detline2 <- matrix(0,4,5) for (i in 1:4) for (j in 1:5){{ sub1 <- mydatadet[mydatadet[,5] ==lvl[i],] Detline2[i,j] <- mean(subset(sub1$lResponse,sub1[,4] == enZ[j])) }} rownames(Detline2) <- lvl colnames(Detline2) <- enZ
graphics.off() windows(width=7,3.5) par(mfrow=c(1,2)) par(mar= c(4,4,2,1)) plot(lResponse ~ EnzymeConc, data = mydatadet , type = 'n', main = 'With detergent', ylim = c(0.5,3.4), xlab = 'Enzyme Concentration (nM)', ylab = 'log10 Response (RU)') for (i in 1:5) { points(lvl,Detline[,i], type = 'o', col = i, pch = 16) } legend('bottomright',enZ, fill = 1:5, horiz=TRUE, cex = 0.8, title = 'Enzyme') text(0.2,3.3, labels = 'A')
plot(lResponse ~ EnzymeConc, data = mydatadet , type = 'n', main = 'Without detergent', ylim = c(0.5,3.4), xlab = 'Enzyme Concentration (nM)', ylab = '') for (i in 1:5) { points(lvl,Detline2[,i], type = 'o', col = i, pch = 16) } text(0.2,3.3, labels = 'B') ## Detergent plots!
########### Make a box plot of response vs addition of detergent and hardness ################
12 | P a g e
352 353 354 355 356 357 358 359 360 361 362 363
Technical University of Denmark Dept of Mathematics and Computer
graphics.off() windows(width=7, height = 2.5) par(mfrow=c(1,2)) par(mar=c(2.5,4.5,0.8,1)) plot(lResponse ~ DetStock, data = mydata, ylab = 'log10 Response (RU)') text(mydata$DetStock[1],1,labels = 'A') plot(lResponse ~ CaStock, data = mydata, xlab = 'Hardness', ylab = '') text(mydata$CaStock[1],1, labels ='B', pos = 4, offset=2)
13 | P a g e
Você também pode gostar
- Modeling and Parameter Estimation of A Nonlinear Switching Systemin Fed-Batch Culture With PH FeedbackDocumento11 páginasModeling and Parameter Estimation of A Nonlinear Switching Systemin Fed-Batch Culture With PH FeedbackDeepak KumarAinda não há avaliações
- Parametric Tests Guide by Dr. SISWANTODocumento34 páginasParametric Tests Guide by Dr. SISWANTOadindapramitrapsAinda não há avaliações
- PH and Salt Effects On Chiral Separations Using Affinity UltrafiltrationDocumento6 páginasPH and Salt Effects On Chiral Separations Using Affinity UltrafiltrationJosé CatalánAinda não há avaliações
- Thesis OptimaDocumento7 páginasThesis Optimagyuusthig100% (1)
- Chemical Kinetics in Biology: GoalsDocumento28 páginasChemical Kinetics in Biology: GoalsDrJayasimha GoudAinda não há avaliações
- Chemical Kinetics in Biology - Analyzing Reaction RatesDocumento69 páginasChemical Kinetics in Biology - Analyzing Reaction RatesDaniel YakubovichAinda não há avaliações
- Estimation of Serum Creatinine by Routine Jaffes Method and by Dry Chemistry in Icteric and Hemolytic Serum SamplesDocumento8 páginasEstimation of Serum Creatinine by Routine Jaffes Method and by Dry Chemistry in Icteric and Hemolytic Serum Samplessyedamasoomazahra9Ainda não há avaliações
- Optimizing Diafiltration for High-Concentration Protein FormulationsDocumento4 páginasOptimizing Diafiltration for High-Concentration Protein FormulationsVenkata Suryanarayana GorleAinda não há avaliações
- 1040 Method Development and Evaluation PDFDocumento3 páginas1040 Method Development and Evaluation PDFTeresa JeriAinda não há avaliações
- Synthesis of Isoamyl AcetateDocumento10 páginasSynthesis of Isoamyl AcetateMikaila Denise LoanzonAinda não há avaliações
- Parmeter Identification in Dynamical Models of Anaerobic Waste Water TreatmentDocumento14 páginasParmeter Identification in Dynamical Models of Anaerobic Waste Water TreatmentSergio ArangoAinda não há avaliações
- Enzyme KineticsDocumento28 páginasEnzyme KineticsJed Dumadag DanoAinda não há avaliações
- Mathematical Modeling of Biosensors: Enzyme-Substrate Interaction and Biomolecular InteractionDocumento15 páginasMathematical Modeling of Biosensors: Enzyme-Substrate Interaction and Biomolecular InteractionElbahi DjaalabAinda não há avaliações
- ConductometryDocumento19 páginasConductometryAinin ZahratunAinda não há avaliações
- Rodriguez Et Al (2008) - Determination of The Adequate MinimumDocumento9 páginasRodriguez Et Al (2008) - Determination of The Adequate MinimumDaniela De Araujo SampaioAinda não há avaliações
- Biochemistry Lab Review UNTDocumento4 páginasBiochemistry Lab Review UNTSue WalthersAinda não há avaliações
- Estimation of Figures of Merit Using Univariate Statistics For Quantitative Second-Order Multivariate Curve ResolutionDocumento11 páginasEstimation of Figures of Merit Using Univariate Statistics For Quantitative Second-Order Multivariate Curve ResolutionRijalnur HidayatullahAinda não há avaliações
- Running Head The Effect of Peroxidase Concentration On Reaction Rate 1Documento7 páginasRunning Head The Effect of Peroxidase Concentration On Reaction Rate 1migire kennedyAinda não há avaliações
- 2829 FullDocumento2 páginas2829 FullSulaiman RasyidAinda não há avaliações
- Wide-Angle X-Ray Solution Scattering For Protein-Ligand Binding: Multivariate Curve Resolution With Bayesian Confidence IntervalsDocumento11 páginasWide-Angle X-Ray Solution Scattering For Protein-Ligand Binding: Multivariate Curve Resolution With Bayesian Confidence IntervalsFatima Herranz TrilloAinda não há avaliações
- Preparation of SorbentDocumento8 páginasPreparation of SorbentSebastian Genesis ViduyaAinda não há avaliações
- Is There A Rational Method To Purify Proteins From Expert Systems To ProteomicsDocumento12 páginasIs There A Rational Method To Purify Proteins From Expert Systems To ProteomicsGabriela ConsuegraAinda não há avaliações
- DiacereinDocumento6 páginasDiacereinRikin ShahAinda não há avaliações
- Using Plackett Burman designs to test method robustness for fluoroquinolones in eggsDocumento37 páginasUsing Plackett Burman designs to test method robustness for fluoroquinolones in eggsmaria dulceAinda não há avaliações
- Ehp 1306561 s001Documento8 páginasEhp 1306561 s001Yu WaAinda não há avaliações
- Analytical ChemistryDocumento26 páginasAnalytical ChemistryJohannesAinda não há avaliações
- Statistical Process Control LabDocumento22 páginasStatistical Process Control LabCHUKWU VICTORAinda não há avaliações
- Analytical Platforms and Immunoassay Interference: DR Les Perry Consultant Clinical Scientist Croydon University HospitalDocumento27 páginasAnalytical Platforms and Immunoassay Interference: DR Les Perry Consultant Clinical Scientist Croydon University Hospitalmonday125Ainda não há avaliações
- Basics and Statistics of Analytical BiochemistryDocumento91 páginasBasics and Statistics of Analytical BiochemistryChuquiure L. AngelAinda não há avaliações
- 1040 Method Development and EvaluationDocumento4 páginas1040 Method Development and EvaluationRaposo IreneAinda não há avaliações
- Biology Student Research Investigati ONDocumento16 páginasBiology Student Research Investigati ONAneeta ann abrahamAinda não há avaliações
- Anal Chem 84 (2012) 1474-1482Documento9 páginasAnal Chem 84 (2012) 1474-1482Verónica VanderhoevenAinda não há avaliações
- Enzyme Assays A Practical Approach by Robert Eisenthal, Michael DansonDocumento304 páginasEnzyme Assays A Practical Approach by Robert Eisenthal, Michael DansonSara OchoaAinda não há avaliações
- Kinetics of a Reaction Rate LawDocumento15 páginasKinetics of a Reaction Rate LawSebatHian Santiago60% (5)
- University of Zakho Faculty of Science Department of ChemistryDocumento29 páginasUniversity of Zakho Faculty of Science Department of ChemistryAhmed AmirAinda não há avaliações
- Análisis de Trazas ArtículoDocumento31 páginasAnálisis de Trazas ArtículoOmar BojórquezAinda não há avaliações
- Effect of Synthesis Parameters in Polyaniline: in Uence On Yield and Thermal BehaviorDocumento10 páginasEffect of Synthesis Parameters in Polyaniline: in Uence On Yield and Thermal BehaviorRiezl MartinAinda não há avaliações
- Using Modelica-Matlab For Parameter Estimation in A Bioethanol Fermentation ModelDocumento13 páginasUsing Modelica-Matlab For Parameter Estimation in A Bioethanol Fermentation ModelwariascAinda não há avaliações
- Design of Experiments - Applications 2013Documento120 páginasDesign of Experiments - Applications 2013Daoud IssaAinda não há avaliações
- The Amount of Acetic Acid (ML) To Neutralise Sodium-Bicarbonate Ethanoic-Acid Solution at Different Concentrations (Molar) of Acetic AcidDocumento8 páginasThe Amount of Acetic Acid (ML) To Neutralise Sodium-Bicarbonate Ethanoic-Acid Solution at Different Concentrations (Molar) of Acetic AcideoqcwxhfhsedwcddmyAinda não há avaliações
- Nonlinear Identification of PH Process by Using Nnarx Model: M.Rajalakshmi, G.Saravanakumar, C.KarthikDocumento5 páginasNonlinear Identification of PH Process by Using Nnarx Model: M.Rajalakshmi, G.Saravanakumar, C.KarthikGuru SaravanaAinda não há avaliações
- Design of A Portable Urine Glucose Monitoring System For Health CareDocumento12 páginasDesign of A Portable Urine Glucose Monitoring System For Health CareaksssudhakarAinda não há avaliações
- Lorno HPLCDocumento5 páginasLorno HPLCmostafaAinda não há avaliações
- Biochemsitry 3304 Midterm 3 - KeyDocumento12 páginasBiochemsitry 3304 Midterm 3 - Keyabelopez12Ainda não há avaliações
- Molecularly Imprinted Electrochemical Sensing of Urinary Melatonin in A Microfluidic SystemDocumento11 páginasMolecularly Imprinted Electrochemical Sensing of Urinary Melatonin in A Microfluidic SystemgpaivAinda não há avaliações
- Predicting Protein-Ligand Interactions Using Iterative Stochastic Elimination AlgorithmDocumento149 páginasPredicting Protein-Ligand Interactions Using Iterative Stochastic Elimination AlgorithmbgorelikAinda não há avaliações
- BSC - T1 - 2024 - Assignment - Experiment ReportDocumento25 páginasBSC - T1 - 2024 - Assignment - Experiment ReportrebeccazhoweeAinda não há avaliações
- Journal of Chromatography A: David Pfister, Fabian Steinebach, Massimo MorbidelliDocumento9 páginasJournal of Chromatography A: David Pfister, Fabian Steinebach, Massimo MorbidelliJames EdwardsAinda não há avaliações
- Koushik Ponnuru: Research PlanDocumento19 páginasKoushik Ponnuru: Research PlanKoushik PonnuruAinda não há avaliações
- General Concepts and Experimental ErrorDocumento45 páginasGeneral Concepts and Experimental Errorboygenius21_39464798Ainda não há avaliações
- Faculty of Applied Sciences: Please Tick Lab Report Assignment ProjectDocumento3 páginasFaculty of Applied Sciences: Please Tick Lab Report Assignment Projectlim wei yangAinda não há avaliações
- Reverse-Phase High-Performance Liquid Chromatography Analysis of BiomoleculesDocumento47 páginasReverse-Phase High-Performance Liquid Chromatography Analysis of Biomoleculesabebaw matebuAinda não há avaliações
- CN2102 Studio Lab Report 3Documento4 páginasCN2102 Studio Lab Report 3prisharameshbabu5Ainda não há avaliações
- BSC - T1 - 2024 - Assignment - Experiment Report (Final)Documento25 páginasBSC - T1 - 2024 - Assignment - Experiment Report (Final)rebeccazhoweeAinda não há avaliações
- 2016 ECCMID Poster CMV Validation ELITe IngeniusDocumento1 página2016 ECCMID Poster CMV Validation ELITe IngeniusDVAinda não há avaliações
- Fuhrer 2011Documento7 páginasFuhrer 2011CoAinda não há avaliações
- Clinical ChemistryDocumento19 páginasClinical Chemistrythrowawy100% (2)
- Instrumental Analysis Lecture Notes IIDocumento56 páginasInstrumental Analysis Lecture Notes IIcoolhemakumar100% (3)
- Thermometric Titrimetry: International Series of Monographs in Analytical ChemistryNo EverandThermometric Titrimetry: International Series of Monographs in Analytical ChemistryAinda não há avaliações
- Thermodynamics Experiment Number 3 Radiation Correction Sarthak PDFDocumento9 páginasThermodynamics Experiment Number 3 Radiation Correction Sarthak PDFSarthak JainAinda não há avaliações
- Problem 2C.4Documento9 páginasProblem 2C.4NajwaAinayaZawaidAinda não há avaliações
- Modeling of Heat Transfer in Two-Phase Flow Using The Level-Set MethodDocumento125 páginasModeling of Heat Transfer in Two-Phase Flow Using The Level-Set Methodnaru_saAinda não há avaliações
- Measures of Central Tendency - FinalDocumento33 páginasMeasures of Central Tendency - FinalPranav KumarAinda não há avaliações
- Basic Calculus-Q3-Module-2Documento23 páginasBasic Calculus-Q3-Module-2macgigaonlinestore100% (1)
- Intersection A Curve and A LineDocumento16 páginasIntersection A Curve and A LineIr HeriawanAinda não há avaliações
- GATE Electrical Engineering GuideDocumento3 páginasGATE Electrical Engineering Guidesampurnanand mishraAinda não há avaliações
- Oup TransitionDocumento4 páginasOup TransitionBjorn AbuboAinda não há avaliações
- Theis Confined Aquifer Test AnalysisDocumento5 páginasTheis Confined Aquifer Test AnalysisAlberto RicardoAinda não há avaliações
- Transformer and Induction Motor Concepts Explained in DetailDocumento2 páginasTransformer and Induction Motor Concepts Explained in Detailshiny_vikram6365Ainda não há avaliações
- Lesson Plan 2 Complmentary and Supplementary AnglesDocumento5 páginasLesson Plan 2 Complmentary and Supplementary AnglesCristine Castulo PreguntaAinda não há avaliações
- A Student Will Lead A PrayerDocumento11 páginasA Student Will Lead A PrayerEsther Joy Perez100% (3)
- Birational Geometry Caucher Birkar PDFDocumento35 páginasBirational Geometry Caucher Birkar PDFhuangrj9Ainda não há avaliações
- Support axial loads with thrust bearingsDocumento18 páginasSupport axial loads with thrust bearingsReyAinda não há avaliações
- RX3i CPU Reference ManualDocumento589 páginasRX3i CPU Reference ManualEn FaizulAinda não há avaliações
- Game Theory1Documento15 páginasGame Theory1Vishwas tomarAinda não há avaliações
- 15 Traveling Salesman Problem: AuthorDocumento25 páginas15 Traveling Salesman Problem: AuthorRama Krishna MuttavarapuAinda não há avaliações
- Dissolution: How To Calculate Dissolution Calculation in Excel SheetDocumento6 páginasDissolution: How To Calculate Dissolution Calculation in Excel SheetSagar kishor savaleAinda não há avaliações
- Stress and Strain in Dental MaterialsDocumento26 páginasStress and Strain in Dental MaterialsDrSweta GandhiAinda não há avaliações
- Droplet Evaporation With Reference To The Effectiveness of Water Mist CoolingDocumento9 páginasDroplet Evaporation With Reference To The Effectiveness of Water Mist CoolingWhidas PrihantoroAinda não há avaliações
- Newton's Second Law AnalysisDocumento2 páginasNewton's Second Law AnalysisJamiel CatapangAinda não há avaliações
- Calculus For BusinessDocumento4 páginasCalculus For BusinessCherifa FarhatAinda não há avaliações
- Practice Problem Solutions CIDocumento3 páginasPractice Problem Solutions CIatiq mengalAinda não há avaliações
- Lagrangian Dynamics HomeworkDocumento10 páginasLagrangian Dynamics HomeworkAnyiAinda não há avaliações
- An Analysis of The Impacts of Heterogeneity Losses Due To Upscaling in Well Testing SimulationsDocumento127 páginasAn Analysis of The Impacts of Heterogeneity Losses Due To Upscaling in Well Testing SimulationsBreno TavaresAinda não há avaliações
- Algoritmos DistribuidosDocumento221 páginasAlgoritmos DistribuidosDiego LozaAinda não há avaliações
- Experimental Design ResearchDocumento25 páginasExperimental Design ResearchRaghuram SeshabhattarAinda não há avaliações
- NSAIS ROW 2019 ProceedingsDocumento54 páginasNSAIS ROW 2019 ProceedingsmikaelcollanAinda não há avaliações
- Level Order Traversal Using Recursion in C++Documento4 páginasLevel Order Traversal Using Recursion in C++kartik mishraAinda não há avaliações
- TrinMaC Virtual Winter 2022 HS ProblemsDocumento5 páginasTrinMaC Virtual Winter 2022 HS ProblemsbogdanAinda não há avaliações