Escolar Documentos
Profissional Documentos
Cultura Documentos
Microarrays: The Use of Oligonucleotides and cDNA For The Analysis of Gene Expression
Enviado por
vongoclinhgiangTítulo original
Direitos autorais
Formatos disponíveis
Compartilhar este documento
Compartilhar ou incorporar documento
Você considera este documento útil?
Este conteúdo é inapropriado?
Denunciar este documentoDireitos autorais:
Formatos disponíveis
Microarrays: The Use of Oligonucleotides and cDNA For The Analysis of Gene Expression
Enviado por
vongoclinhgiangDireitos autorais:
Formatos disponíveis
reviews
research focus
DDT Vol. 8, No. 3 February 2003
Microarrays: the use of oligonucleotides and cDNA for the analysis of gene expression
J. Carl Barrett and Ernest S. Kawasaki
Completion of the human genome sequence has made it possible to study the expression of the entire human gene complement (>30,000 estimated genes). Aiding in this remarkable feat, DNA microarrays have become the main technological workhorse for gene expression studies. To date, detection platforms for most microarrays have relied on short (25 base) oligonucleotides synthesized in situ, or longer, highly variable length DNAs from PCR amplification of cDNA libraries. A third choice, long (5080 base) oligonucleotide arrays, is now available and might eventually eliminate the use of cDNA arrays. The technology has advanced to such a point that researchers now demand microarrays that are cost-effective and have flexibility and quality assurance. Short- and long-oligonucleotide technologies offer such advantages, and could possibly become the major competing platform in the near future.
w The millennium of 2000 has ushered in
what could be considered as a new era for research into the molecular biology of humans. In early 2000, two research groups the International Human Genome Sequencing Consortium and Celera announced the sequencing of the entire human genome, comprising >3 billion base-pairs of nucleotides. The results of their studies were published simultaneously in the journals Science and Nature [1,2], and indicated that the number of genes in the human genome was much smaller than expected, that is, in the order of 30,00040,000 genes compared with the original estimate of ~100,000 genes. This smaller number of genes was disappointing to those who believed humanity to be top of the evolutionary heap, especially because the genome of the modest nematode Caenorhabditis elegans was found to encode 19,000 genes [3], and the rice (Oryza sativa) genome topped the poll with up to 50,000 genes [4,5].
Historical perspective
This wealth of genomic information has enabled researchers to begin the study of the expression and function of every gene in the human body. But how does one begin to analyze the biology of >30,000 genes simultaneously in one experiment? One answer lies in microarray technology, which, although introduced in the 1990s, has only recently come into widespread use. Although the first gene expression microarray article was published in 1995 [6], the concept of miniaturized ligand-binding assays was developed in the mid- to late-1980s [7]. For example, microspot fluorescent immunoassays were described by Roger Ekins in 1989 [8], and the use of doublefluorescent labels for measuring protein analyte concentration was reviewed by the same author in 1989 [9]. Although not exactly equivalent, the use of fluorescent tags to measure nucleic acid hybridization is the mainstay of most array studies, and is a logical extension of earlier immunoassay studies. Thus, conceptually, the first microarrays were of the antigenantibody type, an area of research that is also beginning to expand rapidly [1014].
J. Carl Barrett Center for Cancer Research National Cancer Institute 31 Center Drive, MSC-2440 Bethesda MD 20892, USA Ernest S. Kawasaki* NCI Advanced Technology Center 8717 Grovemont Circle Gaithersburg MD 20877, USA *tel: +1 301 435 2891 fax: +1 301 402 3134 e-mail: kawasake@mail.nih.gov
Technology overview
The microarray field is a good example of the assembly and convergence of several technologies, including automated DNA sequencing, DNA amplification by PCR, highly efficient oligonucleotide synthesis, nucleic acid labeling chemistries, and bioinformatics. A microarray can be thought of as a miniaturized genehybridization or -detection assay. Instead of measuring signals in assays at the macro level, such as in microtiter plates, membrane blots and test tubes, individual microarray assays
134
www.drugdiscoverytoday.com
1359-6446/02/$ see front matter Published by Elsevier Science Ltd. PII: S1359-6446(02)02578-3
DDT Vol. 8, No. 3 February 2003
research focus
reviews
or elements are measured in microns. For example, such micro-elements range from 20 to 200 m, as opposed to 500010,000 m for the bottomsurface dimensions of microtiter plates. One platform used for printing microarrays is the common microscope slide, with dimensions of 25 mm 75 mm. The latest robotic printers can easily fit 50,000 spots or elements onto a slide if the spots are 100 m in diameter and spaced 50 m apart. Thus, it is possible to place the entire human gene complement on one slide a remarkable accomplishment. Each of the elements contains a DNA sequence from one gene, and is used to measure the expression of its corresponding mRNA (mRNA) in a cell or tissue sample. Messenger RNA from control and experimental samples are used to synthesize fluorescently labeled cDNA or RNA probes, which are then hybridized to the microarray elements. The fluorescent signal of the hybridized probes is measured with a laser scanner capable of detecting emission from a variety of fluorescent dyes. The intensity of the signals from control and experimental samples are directly correlated with the original concentration of mRNA in the cell or tissue, and can therefore be used to deduce whether the expression of a particular gene is upregulated, downregulated, unchanged or absent. The sensitivity of the assay is high; microarrays have been reported to detect the presence of one mRNA per cell, that is, a concentration of one mRNA per 100,000 molecules. This is the basic method by which investigators are now able to study the expression of thousands of genes simultaneously in the cell or tissue type of their choice.
(a)
Library of cDNA clones
(b)
Gene/mRNA sequences Bioinformatics/ oligo design algorithms Perfect match mismatch Bacterial growth Plasmid isolation/ purification PCR amplification/ purification of inserts Printing/fixing or coupling DNA Denaturation/ labeling-hybridization/ washing Laser Scanning/data analysis Scanning/data analysis Photolithographic in situ synthesis * * * oligo set
Labeling-hybridization/ washing Laser
(c)
Gene/mRNA sequences Bioinformatics/oligo design algorithms Oligo synthesis/ purification
Library of Oligos
Printing/fixing or coupling DNA Labeling-hybridization/washing Laser
Scanning/data analysis
Drug Discovery Today
Microarray platforms
The following section describes several microarray platforms used for the study of gene expression. This is not meant to be an exhaustive review of all available
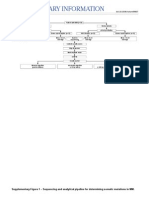
Figure 1. The three microarray platforms. (a) cDNA arrays. A library of cDNA clones contained in 96- or 384-well plates are cultured in bacteria. CDNA inserts from plasmids are amplified and products are spotted onto activated glass slides (25 mm x 75 mm). Probes from mRNA are labeled, hybridized to slides and scanned. (b) Short oligonucleotide arrays. Pairs of oligonucleotides with exact sequence and one mismatch are synthesized in situ. Probes are hybridized to the set that are actually scattered throughout chip and not in one row as shown. (c) Long oligonucleotide arrays. Algorithms are used to design long oligonucleotides of the same size, but with little sequence homology. Protocols for printing, hybridizing and analyses are similar to cDNA arrays.
www.drugdiscoverytoday.com
135
reviews
research focus
DDT Vol. 8, No. 3 February 2003
types, but covers those most relevant to the subject of this article.
cDNA arrays: robotic printing
Before the availability of complete or near-complete eukaryotic genome sequences, genes expressed in cells, tissues or organs were identified through sequence analysis of cDNA banks. cDNA clones from cDNA banks of Arabidopsis and human peripheral blood lymphocytes were used in the construction of the first cDNA expression arrays [6,15]. The inserts of each plasmid were PCR amplified using universal primers corresponding to the vector sequences adjacent to the gene sequences. The amplified products were analyzed by gel electrophoresis, quantitated, arrayed by robotic printing onto surface-modified glass slides, fixed, and then used to study gene expression as described previously (Fig. 1a). Amplified DNAs can be fixed onto the slides electrostatically, or through crosslinking by heat or UV. Alternatively, the PCR products can be covalently attached at their 5 ends to modified slides via an amine or other active group. This method, or minor variations thereof, has been the foundation of most gene expression research where the starting material is plasmids from cDNA banks [1618].
approach for synthesizing oligonucleotides photolithographically using digitally controlled micro-mirrors [2224] could simplify the production of arrays, but the technology is not yet widely available. Further information on similar technologies is available at company websites (febit, http:// www.febit.de; Combimatrix; http://www.combimatrix.com).
Long oligonucleotide arrays: in situ synthesis
The photolithographic method is very efficient at producing thousands or millions of identical arrays, but is somewhat inconvenient and expensive for the creation of new arrays with added or different gene content. Technology has progressed to such a point that publication of a new genome sequence has become commonplace; therefore, the flexibility in designing and producing new arrays to capitalize on this wealth of sequence data is an important issue. The production of new arrays via the synthesis of long, 60-mer oligonucleotides by an ink-jet printing process addresses this point [2527]. In this method, modified ink-jet pumps, similar to those used in printers, are used to dispense 100-picoliter reagent droplets onto a hydrophobic surface containing chemically active hydroxyl groups. The droplets contain phosphoramidite DNA monomers that react and are covalently bound. After washing and de-protection, the process is repeated until the desired oligonucleotide length is reached. The advantages of the in situ inkjet method are that no masks are required, synthesis is faster because each cycle attaches one base (four cycles per base are required with photolithography), and new arrays can be created by simply programming the computer with directions on how to synthesize the new set of oligonucleotide sequences. This versatile system can routinely produce arrays with >25,000 elements [2527].
Short oligonucleotide arrays: in situ synthesis
The first large-scale manufacturing of microarrays was developed by Affymetrix (http://www.affymetrix.com) using photolithographic methods similar to the production of computer chips [19] that were subsequently adapted to gene expression studies [20,21] (Fig. 1b). In brief, synthetic linkers with photolabile protecting groups are attached to a glass substrate, and a mask is used to direct light to predetermined areas on the substrate to remove the exposed groups. These de-protected groups are then available for reaction with bi-functional deoxynucleosides, resulting in chemical coupling. A new mask is used to direct coupling at other sites, and the step is repeated until the desired sequence and length of oligonucleotide is synthesized. The present format is a 1.28 1.28 cm chip containing up to 500,000 different oligonucleotide sequences of 25 bases in length, with each element being 18 18 m in size. Eleven-to-twenty different oligonucleotides are made for each transcript or gene sequence to tile or cover a portion of the 3 end of the mRNA. Oligonucleotides are synthesized as perfect match (PM) and mismatch (MM) pairs [20,21]. The MM oligonucleotide has a one-base mismatch in the center position, and is used as a control to detect background noise and cross-hybridization from unrelated probes. The latest chip design contains the sequences of up to 20,000 genes, but there is space for more genes if fewer oligonucleotides are used to query each gene. A new
Long oligonucleotide arrays: robotic printing
As described previously, the flood of new sequence data enables researchers to design their own arrays in silico, completely bypassing the need to use cumbersome cDNA banks (Fig. 1c). As the cost of manufacturing oligonucleotides has been significantly reduced, obtaining a large gene-library of oligonucleotide sequences is now within the reach of most researchers. Long oligonucleotides are defined here, somewhat arbitrarily, as 4080 bases in length. At this length they can be treated as short cDNAs and there is much less chance of spurious cross-hybridization with unrelated sequences compared with short oligonucleotides. For those experienced in using cDNA arrays, working with long oligonucleotide arrays is a relatively easy transition. The oligonucleotides can be printed onto the same substrates (slides) as cDNAs, with the same printing device,
136
www.drugdiscoverytoday.com
DDT Vol. 8, No. 3 February 2003
research focus
cDNA arrays vs short oligonucleotide arrays NCI cancer cell-line comparisons
reviews
the hybridization and washing conditions are similar, and no new analytical programs for expression analysis are required [28-37]. The design of the oligonucleotides, however, is of upmost importance. Commercial sources have proprietary algorithms for this purpose, although there are several programs in the public domain [3841] if one wishes to create their own set of sequences. Basically, the oligonucleotides should have very similar melting temperatures or GC (guanosinecytosine) content, have very little homology with other oligonucleotides, be entirely contained within an exon, and have no repetitive- or hairpinsequences. Oligonucleotides of any size can be printed onto appropriate substrates, and complex libraries from several commercial sources are now available (ClonTech, http://www.clontech.com; Compugen, http://www.cgen.com; MWG Biotech, http://www.mwg-biotech.com; Operon, http://www.operon.com). This platform is probably the most amenable for researchers who print their own arrays owing to its flexibility, and facile creation of both very high- and low-density arrays.
Comparison of expression experiments
A PubMed query of microarray literature from January 1995 to October 2002 yields a total of >2300 hits. Most of these appeared in the past two years, indicating an exponential increase in the number of microarray publications. The majority of these publications reported the use of short oligonucleotides or cDNAs; only a small number reported the use of long oligonucleotide arrays, as this technology is only now becoming available. Probably owing to cost and time, there have been very few attempts to replicate and/or compare expression data across different platforms. In addition, these types of studies are hindered by many array results not being confirmed by other methods such as northern blots [42] or quantitative RTPCR [43,44]. In addition, there is a general lack of standardized experimental design or controls, so it is almost impossible to compare results across platforms (or even within platforms), and large-scale management and analysis of expression data becomes very difficult [4551]. These problems are being addressed by the Microarray Gene Expression Data Group (MGED group) an international consortium of microarray enthusiasts [52,53]. Their efforts have resulted in the journal Nature [54] having relatively strict guidelines for the submission of microarray data for publication [Minimum Information About a Microarray Experiment (MIAME) guidelines]. Hopefully, other journals will follow suit [55], as such rules will be extremely helpful in removing much of the confusion surrounding the interpretation and replication of microarray data.
Are expression results from different microarray platforms directly comparable? The answer at this point is yes and no, depending on the method of analysis. For example, results published from experiments using Stanford-type cDNA arrays and Affymetrix chips were used to examine the suitability of applying cross-platform data in measuring gene expression [56]. Messenger RNA measurements from 2895 matched genes in 56 cell-lines from the National Cancer Institutes standard panel of 60 cancer cell-lines (NCI 60) were used to compare the two platforms. In general, there was poor correlation between the two platforms in all measurements of similarity, such as clusters of genes and cell-lines, and the study suggested that cDNA array data cannot be combined with the short oligonucleotide array results. As a cautionary note, the authors state that the experiments were carried out at different times, at different laboratories, and using different materials and protocols, and this might explain the discordant results. However, if the results from the two platforms are so dependent on protocol, material and time, as far as the characterization of cancer lines goes, the data could be considered useless. By contrast, the optimist might say the data tell us something about the biology of the cell lines. The authors state that, without a third source of data, they could not recommend one platform above the other.
cDNA vs short oligonucleotide arrays Incyte platform vs Affymetrix
Similar results were obtained with cDNA arrays from Incyte Genomics (http://www.incyte.com) versus the Affymetrix platform in a study of the differences between normal peripheral blood mononuclear cells and large granular lymphocytic leukemia [57]. In this case, experiments were performed in the same laboratory and with the same material, which could explain the poor correlation in the study described previously. However, there were differences in quantitation; for example, the gene encoding perforin showed a 103.0-fold change in the Affymetrix array, and only 3.8-fold change in the cDNA array. Their northern blot analyses gave a value somewhere between the two extremes. Large discrepancies such as these, spread throughout many genes, could easily explain why there seems to be little correlation between the platforms when applying clustering analysis and other algorithms to these data. Another study using cDNA arrays and short oligonucleotide arrays appears to confirm this explanation [58]. The study measured gene expression in a human neuroblastoma cell line, and after treatment of the cells, measurements from Affymetrix chips and Incyte cDNA arrays showed an increase
www.drugdiscoverytoday.com
137
reviews
research focus
DDT Vol. 8, No. 3 February 2003
in the mRNAs of 218 genes and 4 genes, respectively. The four genes were a subset of the 218 RNAs. Nine genes out of the 218 were confirmed as upregulated by RTPCR. In this case, the authors conclude that short oligonucleotide arrays are more reliable for measuring gene expression changes compared with data from cDNA microarrays. In several studies, the short oligonucleotide format appears to have a higher dynamic range than the cDNA format, which could explain why more genes are flagged when studying changes in mRNA levels. However, this is not always the case, as other studies show that the cDNA platforms perform as well or better than the short oligonucleotide arrays [59,60].
samples from primary breast tumors and determined a gene expression signature strongly predictive of poor prognosis, in addition to establishing an expression signature of BRCA1 carriers. These data, along with other publications [2837], indicate that use of long oligonucleotides is an excellent approach to the study of gene expression.
Discussion
The microarray field is now entering a phase of widespread use, similar to the development of PCR technologies in the late 1980s. The availability of complete genomic sequences from many different organisms, advances in microarray instrumentation, and large-scale commercial involvement, gives the researcher a wide choice of platforms from which to choose when conducting gene expression studies. As was the case early in the PCR era, the microarray field is now suffering from enthusiastic use, and growing pains. With several thousand publications already in existence, it has become apparent to many microarray aficionados that a large number of studies cannot be reproduced or replicated, and that related studies from different laboratories, or even in the same laboratory, are not comparable. Establishment of the MIAME guidelines [5255] for publication will go far to alleviate the confusion in the reporting of microarray experiments. These guidelines include details of: (1) how the experiment was designed, (2) the design of the arrays or the name and location of spots on arrays, (3) sample name, extraction and labelling, (4) hybridization protocols, (5) methods for image measurements, and (6) the controls used. Obviously, good experimental design is the first important step, and details of how to properly design microarray experiments are described in recent reviews [50,6264]. There are many sources of variation in microarray studies, and the type of platform used is a major source of this variability. At present, arrays can contain oligonucleotides of 25, 30, 40, 50, 60, 65, 70 to 80 bases in length, and cDNAs of hundreds-of-bases to several kilobases in length. Although some of these platforms are being used for other purposes, such as SNP analysis and diagnostics, it would be helpful if the platforms for gene expression analysis were standardized. The use of short 25-base oligonucleotide and cDNA methods are prevalent, partly because they were the first to be used extensively, and because researchers are usually hesitant to change if a large part of their research programs are based on one particular platform.
Long oligonucleotide array studies
Owing to their relative novelty, there are only a few papers comparing data from long oligonucleotide arrays with other formats. Data from the validation of these types of arrays can be found on the websites of companies producing oligonucleotide libraries (ClonTech, Compugen, MWG Biotech, and Operon) or pre-printed slides (Agilent, http://www.agilent.com). Recently, the assessment of the sensitivity of 50mer arrays compared with cDNA was described [61]. Here, the 50mers and PCR products were spotted on the same slide and their performances evaluated. In this case, no difference in sensitivity between the two were found, and both could detect the equivalent of ~10 mRNA copies per cell. The method was highly specific as cross-hybridization required that the probe have >75% homology with the oligonucleotide. At this level of homology, the probe is probably a related gene and would also have hybridized to its cDNA counterpart. An elegant study comparing short and long oligonucleotide arrays as well as cDNAs was reported by the Rosetta Inpharmatics group (http://www.rosettabio.com) [26]. Oligonucleotides were synthesized in situ with a range of 2060 bases. The study demonstrated that the 60mer length gave the best combination of sensitivity and specificity. The sensitivity was reported as close to one in one-million, or ~0.1 mRNA copies per cell, assuming 100,000 transcripts per cell. This level is as good or better than those reported for short oligonucleotide or cDNA arrays. Comparing the 60mer arrays with cDNA arrays also gave good results. Using an array of one oligonucleotide per gene, they found a close correlation with results from the cDNA array (r = 0.97). That one long oligonucleotide per gene was sufficient for expression studies was an important finding as it significantly simplifies both the in situ synthesis and robotic printing approaches to making arrays. This platform was also used to show that a particular gene expression profile can be used to predict the clinical outcome of breast cancer [27]. The group studied
Advantages and disadvantages Short oligonucleotide arrays
The short oligonucleotide approach has been commercially available for several years and has a strong manufacturing
138
www.drugdiscoverytoday.com
DDT Vol. 8, No. 3 February 2003
research focus
reviews
base, resulting in reagents and chips that are readily available. In addition, there are a large number of published studies using this system, and it is a generally accepted method for expression analysis by many laboratories and journals. With complex eukaryotic genomes, there might be problems with cross-hybridization of the oligonucleotides to unrelated probes because of the short length of the target, but the use of several oligonucleotides per gene appears to help eliminate this problem. One main disadvantage with this approach is that changes are not readily accommodated, for example, when new sequence information becomes available or if different arrays are desired.
concentration. This enables more probe molecules to be annealed, and more than makes up for the shorter length relative to the cDNA arrays. Through appropriate design, long oligonucleotides can distinguish between alternatively spliced mRNAs [65,66] something that is not possible with cDNA arrays. Finally, additions, changes or modifications in the arrays are more straightforward compared with photolithographic methods or cDNA arrays. This is a major advantage to the researcher when new sequence information becomes available or when experimental results require more focused arrays.
Concluding remarks cDNA arrays
The cDNA approach has the advantage that no sequence information is necessary before setting up the arrays. The PCR products from cDNA banks can be synthesized using universal primers, and interesting genes can be sequenced after array analysis. The large size of the PCR product is also helpful in enabling stringent hybridization conditions and lowering cross-hybridization of unrelated genes, although closely related gene families will still be able to anneal to some extent. Although PCR production of DNAs for microarrays is not difficult per se, the large number of DNAs required for complete coverage of a complex genome is taxing for most laboratories. To produce, analyze, purify, aliquot and keep track of 20,00040,000 different PCR products is not easy, and is a difficult feat even for commercial sources dedicated to such a task. The microarray field is in great need of standardization to take full advantage of the tremendous wealth of genomic data. Reducing the number of microarray platforms would be a good first step, and the long oligonucleotide approach is a good candidate to replace cDNA arrays, leaving the researcher with two basic array methods for the study of gene expression. Although there are other, non-array methods for analyzing gene expression, such as SAGE [6770], the simplicity of the oligonucleotide approach makes it the most attractive option for the general research community. The final human genome sequence is scheduled to be released in the spring of 2003, in time for the 50th anniversary of the publication of the structure of DNA. For those engaged in drug development, this means, in essence, that every possible drug target encoded in the genome will be available for testing. Therapeutic drug discovery will no longer be hampered by a shortage of targets but, rather, hindered by an excess of targets. Microarrays will play an essential role in overcoming this obstacle in both target identification and in the long road of drug discovery and development [7176]. Standardized oligonucleotide array platforms will be crucial in helping the pharmaceutical industry to take full advantage of this genomic treasure chest.
Long oligonucleotide arrays
The completion of numerous genomic sequences and increased efficiency of production has combined to make the use long oligonucleotide (5080 bases) for gene expression studies a highly attractive alternative to short oligonucleotide or cDNA arrays. There are several advantages to this platform over the others. Their longer length enables more specificity in hybridization than shorter lengths, and they can therefore be treated similarly to cDNA arrays. Every oligonucleotide is of the same length and has almost the same melting temperature and GC content, enabling more consistent hybridization conditions for every gene on the array. Compared with cDNA arrays, it is much easier to deposit the same concentration of DNA per spot with oligonucleotides. The oligonucleotides are single stranded and do not require a denaturation step as with the cDNA format; this also eliminates the problem of re-naturation, which can decrease hybridization efficiency. The molar concentration of oligonucleotides in arrays of long oligonucleotides is much higher than the short oligonucleotide or cDNA versions, with an ~100-fold difference in
References
1 Venter, J.C. et al. (2001) The sequence of the human genome. Science 291, 13031351 2 Lander, E.S. et al. (2001) Initial sequencing and analysis of the human genome. Nature 409, 860921 3 The C. elegans Sequencing Consortium (1998) Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 20122018 4 Yu, J. et al. (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. Indica). Science 296, 7992 5 Goff, S.A. et al. (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. Japonica). Science 296, 92100 6 Schena, M. et al. (1995) Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467470 7 Ekins, R. and Chu, F.W. (1999) Microarrays: their origins and applications. Trends Biotechnol. 17, 217218
www.drugdiscoverytoday.com
139
reviews
research focus
DDT Vol. 8, No. 3 February 2003
8 Ekins, R. et al. (1989) High specific activity chemiluminescent and fluorescent markers: their potential application to high sensitivity and multi-analyte immunoassays. J. Biolumin. Chemilumin. 4, 5978 9 Ekins, R.P. (1989) Multi-analyte immunoassay. J. Pharm. Biomed. Anal. 7, 155168 10 Hayward, R.E. et al. (2001) Probing the proteome protein arrays and their applications. Drug Discov. Today 6, 12631264 11 Mitchell, P. (2002) A perspective on protein microarrays. Nat. Biotechnol. 20, 225229 12 Templin, M.F. et al. (2002) Protein microarray technology. Trends Biotechnol. 20, 160166 13 Petricoin, E.F. et al. (2002) Clinical proteomics: translating benchside promise into bedside reality. Nat. Rev. Drug Discov. 1, 683695 14 MacBeath, G. (2002) Protein arrays and proteomics. Nat. Genet. 32 (Suppl.), 526532 15 Schena, M. et al. (1996) Parallel human genome analysis: microarraybased expression monitoring of 1000 genes. Proc. Natl. Acad. Sci. U. S. A. 93, 1061410619 16 Duggan, D.J. et al. (1999) Expression profiling using cDNA microarrays. Nat. Genet. 21 (Suppl), 1014 17 Winzeler, E.A. et al. (1999) Fluorescence-based expression monitoring using microarrays. Methods Enzymol. 306, 318 18 Lou, X.J. et al. (2001) Expression monitoring using cDNA microarrays. Methods Mol. Biol. 175, 323340 19 Fodor, S.P.A. et al. (1991) Light-directed, spatially addressable parallel chemical synthesis. Science 251, 767773 20 Lockhart, D.J. et al. (1996) Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat. Biotechnol. 14, 16751680 21 Lipshutz, R.J. et al. (1999) High density synthetic oligonucleotide arrays. Nat. Genet. 21 (Suppl), 2024 22 Singh-Gasson, S. et al. (1999) Maskless fabrication of light-directed oligonucleotide microarrays using a digital micromirror array. Nat. Biotechnol. 17, 974978 23 Cerrina, F. et al. (2002) Biological lithography: development of a maskless microarray synthesizer for DNA chips. Microelectron. Eng. 61-62, 3340 24 Nuwaysir, E.F. et al. (2002) Gene expression analysis using oligonucleotide arrays produced by maskless photolithography. Genome Res. 12, 17491755 25 Blanchard, A.P. et al. (1996) High-density oligonucleotide arrays. Biosens. Bioelectron. 11, 687690 26 Hughes, T.R. et al. (2001) Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat. Biotechnol. 19, 342347 27 van Veer, A. et al. (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530536 28 Chambers, J. et al. (1999) DNA microarrays of the complex human cytomegalovirus genome: profiling kinetic class with drug sensitivity of viral gene expression. J. Virol. 73, 57575766 29 Beaucage, S.L. (2001) Strategies in the preparation of DNA oligonucleotide arrays for diagnostic applications. Curr. Med. Chem. 8, 12131244 30 Dudley, A.M. et al. (2002) Measuring absolute expression with microarrays with a calibrated reference sample and an extended signal intensity range. Proc. Natl. Acad. Sci. U. S. A. 99, 75547559 31 Relogio, A. et al. (2002) Optimization of oligonucleotide-based DNA microarrays. Nucleic Acids Res. 30, e51 32 El Atifi, M. et al. (2002) Long oligonucleotide arrays on nylon for large-scale gene expression analysis. BioTechniques 33, 612618 33 Wang, R-F. et al. (2002) Design and evaluation of oligonucleotidemicroarray method for the detection of human intestinal bacteria in fecal samples. FEMS Microbiol. Lett. 213, 175182 34 Wagner, E.K. et al. (2002) Practical approaches to long oligonucleotidebased DNA microarray: lessons from Herpesviruses. Prog. Nucleic Acid Res. Mol. Biol. 71, 445491 35 Stingley, S.W. et al. (2000) Global analysis of herpes simplex virus type I
36
37 38 39 40
41 42
43 44
45 46 47 48 49
50 51 52
53 54 55 56 57 58 59 60
61 62 63 64
transcription using an oligonucleotide-based DNA microarray. J. Virol. 74, 99169927 Yang, W.C. et al. (2002) General and specific alterations in programming of global viral gene expression during infection by VP16 activation-deficient mutants of herpes simplex virus type I. J. Virol. 76, 1275812774 Wang, D. et al. (2002) Microarray-based detection and genotyping of viral pathogens. Proc. Natl. Acad. Sci. U. S. A. 99, 1568715692 Li, F. and Stormo, G.D. (2001) Selection of optimal DNA oligos for gene expression arrays. Bioinformatics 17, 10671076 Rouillard, J-M. et al. (2002) OligoArray: genome-scale oligonucleotide design for microarrays. Bioinformatics 18, 486487 Chen, H. and Sharp, B.M. (2002) Oliz, a suite of Perl scripts that assist in the design of microarrays using 50mer oligonucleotides from the 3 untranslated region. BMC Bioinformatics 3, 27 Wright, M.A. and Church, G.M. (2002) An open-source oligomicroarray standard for human and mouse. Nat. Biotechnol. 20, 10821083 Taniguchi, M. et al. (2001) Quantitative assessment of DNA microarrays comparison with Northern blot analyses. Genomics 71, 3439 OConnell, J. (2002) RT-PCR in biomedicine. Opportunities arising from the new accessibility of mRNA. Methods Mol. Biol. 193, 317 Bustin, S.A. (2000) Absolute quantitation of mRNA using real-time reverse transcription polymerase chain reaction assays. J. Mol. Endocrinol. 25, 169193 Aach, J. et al. (2000) Systematic management and analysis of yeast gene expression data. Genome Res. 10, 431445 Quackenbush, J. (2001) Computational analysis of microarray data. Nat. Rev. Genet. 2, 418427 Nadon, R. and Shoemaker, J. (2002) Statistical issue with microarrays: processing and analysis. Trends Genet. 18, 265271 Brazma, A. et al. (2002) Microarray data representation, annotation and storage. Adv. Biochem. Eng. Biotechnol. 77, 113139 Holloway, A.J. et al. (2002) Options available from start to finish for obtaining data from DNA microarrays II. Nat. Genet. 32 (Suppl.), 481489 Churchill, G.A. (2002) Fundamentals of experimental design for cDNA microarrays. Nat. Genet. 32 (Suppl.), 490495 Quackenbush, J. (2002) Microarray data normalization and transformation. Nat. Genet. 32 (Suppl.), 496501 Brazma, A. et al. (2001) Minimum information about a microarray experiment (MIAME) towards standards for microarray data. Nat. Genet. 29, 365371 Stoeckert, C.J., Jr et al. (2002) Microarray databases: standards and ontologies. Nat. Genet. 32 (Suppl.), 469473 No author (2002) Microarray standards at last. Nature 419, 323 Ball, C. et al. (2002) Standards for microarray data. Science 298, 539 Kuo, W.P. et al. (2002) Analysis of matched mRNA measurements from two different microarray technologies. Bioinformatics 18, 405412 Kothapalli, R. et al. (2002) Microarray results: how accurate are they? BMC Bioinformatics 3, 22 Li, J. et al. (2002) Differential gene expression patterns revealed by oligonucleotide versus long cDNA arrays. Toxicol. Sci. 69, 383390 Yuen, T. et al. (2002) Accuracy and calibration of commercial oligonucleotide and custom cDNA arrays. Nucleic Acid Res. 30, e48 Rhodes, D.R. et al. (2002) Meta-analysis of microarrays: interstudy validation of gene expression profiles reveals pathway disregulation in prostate cancer. Cancer Res. 62, 44274433 Kane, M.D. et al. (2000) Assessment of the sensitivity and specificity of oligonucleotide (50mer) microarrays. Nucleic Acid Res. 28, 45524557 Kerr, M.K. and Churchill, G.A. (2001) Experimental design for gene expression microarrays. Biostatistics 2, 183201 Yang, Y.H. and Speed, T. (2002) Design issues for cDNA microarray experiments. Nat. Rev. Genet. 3, 579588 Simon, R. et al. (2002) Design studies using DNA microarrays. Genet. Epidemiol. 23, 2136
140
www.drugdiscoverytoday.com
DDT Vol. 8, No. 3 February 2003
research focus
reviews
65 Clark, T.A. et al. (2002) Genomewide analysis of mRNA processing in yeast using splicing-specific microarrays. Science 296, 907910 66 Maniatis, T. and Tasic, B. (2002) Alternative pre-mRNA splicing and proteome expansion in metazoans. Nature 418, 236243 67 Liang, P. (2002) SAGE Genie: a suite with panoramic view of gene expression. Proc. Natl. Acad. Sci. U. S. A. 99, 1154711548 68 Evans, S.J. et al. (2002) Evaluation of Affymetrix gene chip sensitivity in rat hippocampal tissue using SAGE analysis. Eur. J. Neurosci. 16, 409413 69 Patino, W.D. et al. (2002) Serial analysis of gene expression: technical considerations and applications to cardiovascular biology. Circ. Res. 91, 565569 70 Soulet, D. and Rivest, S. (2002) Perspective: how to make microarray, serial analysis of gene expression, and proteomic relevant to day-to-day endocrine problems and physiological systems. Endocrinololgy 143, 19952001
71 Somogyi, R. and Greller, L.D. (2001) The dynamics of molecular networks: applications to therapeutic discovery. Drug Discov. Today 6, 12671277 72 Clarke, P.A. et al. (2001) Gene expression microarray analysis in cancer biology, pharmacology, and drug development: progress and potential. Biochem. Pharmacol. 62, 13111336 73 Ulrich, R. and Friend, S.H. (2002) Toxigenomics and drug discovery: will new technologies help us produce better drugs? Nat. Rev. Drug Discov. 1, 8488 74 Hamadeh, H.K. et al. (2002) Gene expression analysis reveals chemical-specific profiles. Toxicol. Sci. 67, 219231 75 Petricoin, E.F., III et al. (2002) Medical applications of microarray technologies: a regulatory science perspective. Nat. Genet. 32 (Suppl.), 474479 76 Gerhold, D.L. et al. (2002) Better therapeutics through microarrays. Nat. Genet. 32 (Suppl.), 547552
Drug Discovery Today will protect your identity!
Here is an unrivalled opportunity to get off your chest those things that really irritate you! and to be able to tell the relevant people what really irritates you without them knowing it is you! Send in your letters and get some real debate going! Please send all contributions to Dr Joanna Owens e-mail: joanna.owens@elsevier.com
Publication of letters is subject to editorial discretion
Contributions to Monitor
We welcome recommendations of papers for review within Monitor, in the fields of combinatorial chemistry, pharmacogenomics, pharmacoproteomics, bioinformatics, new therapeutic targets, high throughput screening, new drug delivery technologies and other promising lines of research. Details of recent papers or those in press should be directed to Dr Steve Carney, Editor, Drug Discovery Today, Elsevier Science London, 84 Theobalds Road, London, UK WC1X 8RR. tel: +44 207 611 4132, fax: +44 207 611 4485, e-mail: s.carney@elsevier.com
Contributions to Profiles
We welcome contributions for the Profiles series, which gives a commentary on promising lines of research, new technologies and progress in therapeutic areas. Articles should provide an accurate summary of the essential facts together with an expert commentary to provide a perspective. Brief outlines of proposed articles should be directed to the Monitor Editor (see below). Articles for publication in Monitor are subject to peer review and occasionally may be rejected or, as is more often the case, authors may be asked to revise their contribution. The Monitor Editor also reserves the right to edit articles after acceptance. All suggestions or queries relating to Monitor should be addressed to Dr Steve Carney, Editor, Drug Discovery Today, Elsevier Science London, 84 Theobalds Road, London, UK WC1X 8RR. tel: +44 207 611 4132, fax: +44 207 611 4485, e-mail: s.carney@elsevier.com
www.drugdiscoverytoday.com
141
Você também pode gostar
- Minimal Residual Disease Testing: Current Innovations and Future DirectionsNo EverandMinimal Residual Disease Testing: Current Innovations and Future DirectionsTodd E. DruleyAinda não há avaliações
- microtoxDocumento7 páginasmicrotoxtexto.sarlAinda não há avaliações
- Dna Chip Technologies: Seung Yong Hwang and Geunbae LimDocumento5 páginasDna Chip Technologies: Seung Yong Hwang and Geunbae LimPulkit ShrivastavaAinda não há avaliações
- Gene Arrays Reveal Cell FunctionsDocumento15 páginasGene Arrays Reveal Cell FunctionsRizkia Milladina HidayatullohAinda não há avaliações
- Universiti Malaysia Sabah Faculty of Science and Human Resources Hgo7 BiotechnologyDocumento9 páginasUniversiti Malaysia Sabah Faculty of Science and Human Resources Hgo7 BiotechnologyShalini MuthuAinda não há avaliações
- 2023 07 03 547464v1 FullDocumento36 páginas2023 07 03 547464v1 FullasdfsoidjfoiqjwefAinda não há avaliações
- 1 Part2Documento13 páginas1 Part2Ali AFadelAinda não há avaliações
- CFG PBL DivvDocumento6 páginasCFG PBL DivvWhatsoever BlogsAinda não há avaliações
- MicroarrayDocumento15 páginasMicroarrayKedar Ghimire100% (9)
- Microarray TechnologyDocumento4 páginasMicroarray TechnologyKristyn RiveraAinda não há avaliações
- A Concise Guide To cDNA Microarray Analysis - IIDocumento27 páginasA Concise Guide To cDNA Microarray Analysis - IIaaasidAinda não há avaliações
- New Microsoft Office Word DocumentDocumento5 páginasNew Microsoft Office Word DocumentNaga SamanthAinda não há avaliações
- biotechnologyedited-180226164222Documento10 páginasbiotechnologyedited-180226164222kush PrajapatiAinda não há avaliações
- DNA Microarrays: by Mohammed Mashik Kommareddy Krithik Yadu Krishnan Mubashir Deepak A SDocumento30 páginasDNA Microarrays: by Mohammed Mashik Kommareddy Krithik Yadu Krishnan Mubashir Deepak A SVipin GeorgeAinda não há avaliações
- Jaipur National University: Presented By: Richa Kumari Branch: B.Tech Biotech 6 SEMDocumento27 páginasJaipur National University: Presented By: Richa Kumari Branch: B.Tech Biotech 6 SEMRamkrishna100% (1)
- B.tech. Biotechnology NotesDocumento14 páginasB.tech. Biotechnology NotesMudit MisraAinda não há avaliações
- Next Generation SequencingDocumento7 páginasNext Generation SequencingAvani KaushalAinda não há avaliações
- 6456456accepted Manuscript (Woerner)Documento47 páginas6456456accepted Manuscript (Woerner)RodrigoAinda não há avaliações
- Next Generation Sequencing - ReviewDocumento58 páginasNext Generation Sequencing - Reviewgvdt19Ainda não há avaliações
- Micro Array ChipDocumento8 páginasMicro Array ChipMudit MisraAinda não há avaliações
- QuarterlyRevBiophys PDFDocumento33 páginasQuarterlyRevBiophys PDFHenryAinda não há avaliações
- Genechip Technology and Its Applications: September 2005Documento7 páginasGenechip Technology and Its Applications: September 2005Anonymous GGL7k0PWAinda não há avaliações
- Next Generation SequencingDocumento9 páginasNext Generation Sequencinghumorboy123Ainda não há avaliações
- Single Cell Transcriptomics - Methods and Protocols-Humana Press - (Methods in Molecular Biology, 2584) Raffaele A. Calogero, Vladimir Benes - 2022Documento394 páginasSingle Cell Transcriptomics - Methods and Protocols-Humana Press - (Methods in Molecular Biology, 2584) Raffaele A. Calogero, Vladimir Benes - 2022julio castilloAinda não há avaliações
- Transcriptome Profiling: Methods and Applications-A Review: November 2017Documento12 páginasTranscriptome Profiling: Methods and Applications-A Review: November 2017Phelix O DaniyanAinda não há avaliações
- Overview of Next Generation Sequencing TechnologiesDocumento12 páginasOverview of Next Generation Sequencing TechnologiesiuventasAinda não há avaliações
- DNA MicroarrayDocumento17 páginasDNA Microarraygargayush100% (1)
- Diagnóstico Molecular-ENARM 2008: Chapter 62. Principles of Human GeneticsDocumento29 páginasDiagnóstico Molecular-ENARM 2008: Chapter 62. Principles of Human GeneticsEmanuel GonzàlezAinda não há avaliações
- Transcriptome Profiling Methods Review: RNA-Seq, Microarrays, SAGE & MPSSDocumento12 páginasTranscriptome Profiling Methods Review: RNA-Seq, Microarrays, SAGE & MPSSResearch SolutionsAinda não há avaliações
- DNA RecombinationnewDocumento34 páginasDNA Recombinationnewsecretary.kimeniahAinda não há avaliações
- MicrosDocumento8 páginasMicrosduchess juliane mirambelAinda não há avaliações
- Research Paper On Dna MicroarrayDocumento7 páginasResearch Paper On Dna Microarrayafnhbijlzdufjj100% (1)
- Genome Sequencing: Dr. P. Balaji Vysya College, HosurDocumento72 páginasGenome Sequencing: Dr. P. Balaji Vysya College, HosurBalaji PaulrajAinda não há avaliações
- Next-Generation DNA Sequencing MethodsDocumento18 páginasNext-Generation DNA Sequencing MethodsFederico BerraAinda não há avaliações
- DNA Microarray Technology: A Powerful Tool for Gene Expression AnalysisDocumento5 páginasDNA Microarray Technology: A Powerful Tool for Gene Expression AnalysisNoorAinda não há avaliações
- ArticleDocumento8 páginasArticleLe SmithAinda não há avaliações
- Exploring Plant Transcriptomes Using Ultra High-Throughput SequencingDocumento11 páginasExploring Plant Transcriptomes Using Ultra High-Throughput SequencingElisabeth ReyesAinda não há avaliações
- Design Issues For cDNA Microarray ExperimentsDocumento10 páginasDesign Issues For cDNA Microarray ExperimentsRoss CUIAinda não há avaliações
- Transcriptome Profiling Methods and ApplicationsDocumento12 páginasTranscriptome Profiling Methods and ApplicationsKito TongHuiAinda não há avaliações
- Single-Cell Analysis - WikipediaDocumento13 páginasSingle-Cell Analysis - WikipediaEmka HudaAinda não há avaliações
- Tecnologias de Secuenciacion de Siguiente GeneracionDocumento16 páginasTecnologias de Secuenciacion de Siguiente Generacion'Carlos SánchezAinda não há avaliações
- Dnachip1 151213150459Documento27 páginasDnachip1 151213150459125078011Ainda não há avaliações
- Analysis of Gene Expression in Cancer Cell Lines Identifies Candidate Markers For Pancreatic Tumorigenesis and MetastasisDocumento13 páginasAnalysis of Gene Expression in Cancer Cell Lines Identifies Candidate Markers For Pancreatic Tumorigenesis and MetastasisAbhishek BardhanAinda não há avaliações
- Emerging DNA Sequencing Technologies For Human Genomic Medicine PDFDocumento9 páginasEmerging DNA Sequencing Technologies For Human Genomic Medicine PDFGabitza GabiAinda não há avaliações
- Attachment ADocumento10 páginasAttachment Atanpreet_makkadAinda não há avaliações
- Journal of Microbiological MethodsDocumento8 páginasJournal of Microbiological Methodsalxo91Ainda não há avaliações
- Artigo CientíficoDocumento6 páginasArtigo CientíficoMichele CalijorneAinda não há avaliações
- Revised TERM PAPERDocumento19 páginasRevised TERM PAPERVikal RajputAinda não há avaliações
- DNA Microarrays: Applications and TechniquesDocumento29 páginasDNA Microarrays: Applications and TechniquesShubham PrasadAinda não há avaliações
- Dna Microarray Technology: Application in Cancer Gene Detection and DiscoveryDocumento36 páginasDna Microarray Technology: Application in Cancer Gene Detection and DiscoveryOmotoso Toyin-TokiAinda não há avaliações
- Dna Biosensor ThesisDocumento7 páginasDna Biosensor ThesisBuyEssaysTulsa100% (2)
- Shokralla Et Al. 2015Documento7 páginasShokralla Et Al. 2015pdf2006Ainda não há avaliações
- Evolutionary Neural NetworkDocumento19 páginasEvolutionary Neural NetworkchithrasreemodAinda não há avaliações
- Genes: Principles of Genetic EngineeringDocumento21 páginasGenes: Principles of Genetic EngineeringCarljustine SablayAinda não há avaliações
- Sage Technology and Its ApplicationsDocumento61 páginasSage Technology and Its Applicationsthamizh555Ainda não há avaliações
- RibotypingDocumento2 páginasRibotypingRobotrixAinda não há avaliações
- Progress in DNA-Seq GEDocumento80 páginasProgress in DNA-Seq GEPMIB Matrikulasi FKUI 2018/2019Ainda não há avaliações
- DNA Sequencing: MethodsDocumento89 páginasDNA Sequencing: MethodsAshraf SawaftaAinda não há avaliações
- Day 2 General Microarray Lecture - Ver11706Documento48 páginasDay 2 General Microarray Lecture - Ver11706amitAinda não há avaliações
- Comparison of High Throughput Next GenerDocumento12 páginasComparison of High Throughput Next GenerAni IoanaAinda não há avaliações
- Protein Microarray TechnologyDocumento7 páginasProtein Microarray TechnologyvongoclinhgiangAinda não há avaliações
- 6Documento19 páginas6vongoclinhgiangAinda não há avaliações
- 4Documento7 páginas4vongoclinhgiangAinda não há avaliações
- DNA Microarrays Paper 2010Documento25 páginasDNA Microarrays Paper 2010Maya FrostAinda não há avaliações
- 5Documento8 páginas5vongoclinhgiangAinda não há avaliações
- 3Documento8 páginas3vongoclinhgiangAinda não há avaliações
- 4Documento14 páginas4vongoclinhgiangAinda não há avaliações
- Reviews: DNA Microarrays: Translation of The Genome From Laboratory To ClinicDocumento8 páginasReviews: DNA Microarrays: Translation of The Genome From Laboratory To ClinicvongoclinhgiangAinda não há avaliações
- 2Documento11 páginas2vongoclinhgiangAinda não há avaliações
- 1 3Documento6 páginas1 3vongoclinhgiangAinda não há avaliações
- Withanolides of Datura Fastuosa LeavesDocumento3 páginasWithanolides of Datura Fastuosa LeavesvongoclinhgiangAinda não há avaliações
- Biomedical PolymersDocumento470 páginasBiomedical Polymersvongoclinhgiang100% (3)
- Wolstenholme Et Al. (2011)Documento10 páginasWolstenholme Et Al. (2011)DaBid Lopez RodriguezAinda não há avaliações
- Molecular Virology, 2013 Edition (PDF) (DR - Carson) VRG PDFDocumento1.013 páginasMolecular Virology, 2013 Edition (PDF) (DR - Carson) VRG PDFYuridia Rodríguez100% (3)
- BGi RNA-Seq AnalysisDocumento19 páginasBGi RNA-Seq Analysisshashikanth_marriAinda não há avaliações
- Tesis doctoral de Bryan Keith Holland - “Discovery of mature microRNA sequences within the protein-coding regions of global HIV-1 genomes - Predictions of novel mechanisms for viral infection and pathogenicity”Documento183 páginasTesis doctoral de Bryan Keith Holland - “Discovery of mature microRNA sequences within the protein-coding regions of global HIV-1 genomes - Predictions of novel mechanisms for viral infection and pathogenicity”Omar Alejandro Herrera ArenasAinda não há avaliações
- 2ndSemesterLac OPERON1F1 DR KPDocumento23 páginas2ndSemesterLac OPERON1F1 DR KPAlina RafiqueAinda não há avaliações
- BSC Biochemistry Syllabus CBCSDocumento20 páginasBSC Biochemistry Syllabus CBCSRama Krishna SaiAinda não há avaliações
- Test Bank For Human Genetics Concepts and Applications 9th Edition Ricki LewisDocumento14 páginasTest Bank For Human Genetics Concepts and Applications 9th Edition Ricki LewisEthel Carver100% (33)
- Lecture 4 - Transcription and Post Transcription ModificationsDocumento73 páginasLecture 4 - Transcription and Post Transcription ModificationskibzwanjikuAinda não há avaliações
- Mid-Old Cells Are A Potential Target For Anti-Aging Interventions in The ElderlyDocumento16 páginasMid-Old Cells Are A Potential Target For Anti-Aging Interventions in The ElderlyecmelloAinda não há avaliações
- Applied Biosystems 7300 Real-Time PCR SystemDocumento4 páginasApplied Biosystems 7300 Real-Time PCR SystemLiviu Athos TamasAinda não há avaliações
- Wright - Genes and Common Diseases (Cambridge, 2007)Documento561 páginasWright - Genes and Common Diseases (Cambridge, 2007)Mohamed IssamAinda não há avaliações
- Molecular Basis of InheritanceDocumento12 páginasMolecular Basis of InheritanceIshvaryaAinda não há avaliações
- Genetic Code 12Documento16 páginasGenetic Code 12Jahnavi JoshiAinda não há avaliações
- University of Texas BIO 325 ExamDocumento6 páginasUniversity of Texas BIO 325 ExamGeetika InstaAinda não há avaliações
- Combinepdf 7Documento129 páginasCombinepdf 7Joe JosephAinda não há avaliações
- Test Bank For Introduction To Genetic Analysis Eleventh EditionDocumento18 páginasTest Bank For Introduction To Genetic Analysis Eleventh Editiondaonhatc2ddrq100% (25)
- Rajan Et Al., 2013, Galectin-1 Gadus MorhuaDocumento11 páginasRajan Et Al., 2013, Galectin-1 Gadus MorhuaLulu SetiyabudiAinda não há avaliações
- University of Rajshahi MSC SyllabusDocumento16 páginasUniversity of Rajshahi MSC SyllabusProf. Dr. M. KhalequzzamanAinda não há avaliações
- Comparative Population Genomics of Maize Domestication and Improvement Hufford Et Al. 2012Documento6 páginasComparative Population Genomics of Maize Domestication and Improvement Hufford Et Al. 2012Cindi ContrerasAinda não há avaliações
- Banana Housekeeping Reference GenesDocumento14 páginasBanana Housekeeping Reference GenesShuhaidah SalehinAinda não há avaliações
- 8 Mader Biology 11th EdDocumento38 páginas8 Mader Biology 11th EdRJ De JesusAinda não há avaliações
- Genetic Codes With No Dedicated Stop Codon Context Dependent Translation TerminationDocumento13 páginasGenetic Codes With No Dedicated Stop Codon Context Dependent Translation TerminationAarcamAinda não há avaliações
- Efficient Ligation of DNADocumento21 páginasEfficient Ligation of DNAMariaAinda não há avaliações
- NUCLEIC ACID METABOLISM COMPLETE NOTES (B.Pharm 2nd Sem) PDFDocumento16 páginasNUCLEIC ACID METABOLISM COMPLETE NOTES (B.Pharm 2nd Sem) PDFAmit Bansal100% (2)
- Gene Structure Lecture 2Documento36 páginasGene Structure Lecture 2AlexTsuiAinda não há avaliações
- 1 s2.0 S1074761321003654 MainDocumento35 páginas1 s2.0 S1074761321003654 MainIoana BiticaAinda não há avaliações
- Initial Genome Sequencing and Analysis of Multiple Myeloma - SuppDocumento75 páginasInitial Genome Sequencing and Analysis of Multiple Myeloma - SuppShantanu KumarAinda não há avaliações
- Complete Transcription Mechanisms 2020 2021Documento57 páginasComplete Transcription Mechanisms 2020 2021Azer AzonfackAinda não há avaliações
- Effects of Quetiapine On DNA Methylation in Neuroblastoma CellsDocumento5 páginasEffects of Quetiapine On DNA Methylation in Neuroblastoma CellsafpiovesanAinda não há avaliações
- Magnifection-A New Platform For Expressing RecombinantDocumento7 páginasMagnifection-A New Platform For Expressing Recombinantricky_connollyAinda não há avaliações
- Why We Die: The New Science of Aging and the Quest for ImmortalityNo EverandWhy We Die: The New Science of Aging and the Quest for ImmortalityNota: 3.5 de 5 estrelas3.5/5 (2)
- 10% Human: How Your Body's Microbes Hold the Key to Health and HappinessNo Everand10% Human: How Your Body's Microbes Hold the Key to Health and HappinessNota: 4 de 5 estrelas4/5 (33)
- The Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceNo EverandThe Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceNota: 4.5 de 5 estrelas4.5/5 (515)
- Masterminds: Genius, DNA, and the Quest to Rewrite LifeNo EverandMasterminds: Genius, DNA, and the Quest to Rewrite LifeAinda não há avaliações
- The Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindNo EverandThe Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindNota: 4.5 de 5 estrelas4.5/5 (93)
- Crypt: Life, Death and Disease in the Middle Ages and BeyondNo EverandCrypt: Life, Death and Disease in the Middle Ages and BeyondNota: 4 de 5 estrelas4/5 (3)
- When the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisNo EverandWhen the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisNota: 3.5 de 5 estrelas3.5/5 (2)
- This Is Your Brain On Parasites: How Tiny Creatures Manipulate Our Behavior and Shape SocietyNo EverandThis Is Your Brain On Parasites: How Tiny Creatures Manipulate Our Behavior and Shape SocietyNota: 3.5 de 5 estrelas3.5/5 (31)
- The Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionNo EverandThe Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionNota: 4 de 5 estrelas4/5 (811)
- All That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesNo EverandAll That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesNota: 4.5 de 5 estrelas4.5/5 (397)
- The Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorNo EverandThe Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorAinda não há avaliações
- Wayfinding: The Science and Mystery of How Humans Navigate the WorldNo EverandWayfinding: The Science and Mystery of How Humans Navigate the WorldNota: 4.5 de 5 estrelas4.5/5 (18)
- Undeniable: How Biology Confirms Our Intuition That Life Is DesignedNo EverandUndeniable: How Biology Confirms Our Intuition That Life Is DesignedNota: 4 de 5 estrelas4/5 (11)
- The Lives of Bees: The Untold Story of the Honey Bee in the WildNo EverandThe Lives of Bees: The Untold Story of the Honey Bee in the WildNota: 4.5 de 5 estrelas4.5/5 (44)
- The Second Brain: A Groundbreaking New Understanding of Nervous Disorders of the Stomach and IntestineNo EverandThe Second Brain: A Groundbreaking New Understanding of Nervous Disorders of the Stomach and IntestineNota: 4 de 5 estrelas4/5 (17)
- Eels: An Exploration, from New Zealand to the Sargasso, of the World's Most Mysterious FishNo EverandEels: An Exploration, from New Zealand to the Sargasso, of the World's Most Mysterious FishNota: 4 de 5 estrelas4/5 (30)
- A Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsNo EverandA Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsNota: 4.5 de 5 estrelas4.5/5 (4)
- Human Errors: A Panorama of Our Glitches, from Pointless Bones to Broken GenesNo EverandHuman Errors: A Panorama of Our Glitches, from Pointless Bones to Broken GenesNota: 3.5 de 5 estrelas3.5/5 (56)
- The Mind & The Brain: Neuroplasticity and the Power of Mental ForceNo EverandThe Mind & The Brain: Neuroplasticity and the Power of Mental ForceAinda não há avaliações
- Gathering Moss: A Natural and Cultural History of MossesNo EverandGathering Moss: A Natural and Cultural History of MossesNota: 4.5 de 5 estrelas4.5/5 (347)
- The Dog Who Couldn't Stop Loving: How Dogs Have Captured Our Hearts for Thousands of YearsNo EverandThe Dog Who Couldn't Stop Loving: How Dogs Have Captured Our Hearts for Thousands of YearsAinda não há avaliações
- Younger for Life: Feel Great and Look Your Best with the New Science of AutojuvenationNo EverandYounger for Life: Feel Great and Look Your Best with the New Science of AutojuvenationNota: 4 de 5 estrelas4/5 (1)
- Darwin's Dangerous Idea: Evolution and the Meaning of LifeNo EverandDarwin's Dangerous Idea: Evolution and the Meaning of LifeNota: 4 de 5 estrelas4/5 (523)
- Why We Sleep: Unlocking the Power of Sleep and DreamsNo EverandWhy We Sleep: Unlocking the Power of Sleep and DreamsNota: 4.5 de 5 estrelas4.5/5 (2083)
- Lymph & Longevity: The Untapped Secret to HealthNo EverandLymph & Longevity: The Untapped Secret to HealthNota: 4.5 de 5 estrelas4.5/5 (13)