Escolar Documentos
Profissional Documentos
Cultura Documentos
D Dipole
Enviado por
ogangurelDescrição original:
Título original
Direitos autorais
Formatos disponíveis
Compartilhar este documento
Compartilhar ou incorporar documento
Você considera este documento útil?
Este conteúdo é inapropriado?
Denunciar este documentoDireitos autorais:
Formatos disponíveis
D Dipole
Enviado por
ogangurelDireitos autorais:
Formatos disponíveis
Predicting quaternary protein structure
Dimer 2-fold axes are perpendicular to monomer dipole
moments
Ogan Gurel and Prof. Wayne A. Hendrickson
6 March 1996
One of the great unsolved problems in modern biology is the protein folding problem: namely, the
prediction of the threedimensional structure of proteins from their linear amino acid sequence. Most study
in this area has focussed on secondary and tertiary structure prediction with relatively little emphasis on
quaternary structure. In general biophysical terms, it is felt that secondary structural elements such as α
helices and βsheets largely depend upon the minimization of van der Waals steric contacts while for
tertiary structure the hydrophobic effect is a major
determining factor. This work proposes that electrostatic
interactions — in particular, the intermolecular alignment of
protein subunit dipole moments — accounts for (and can
predict) the symmetry elements that define a protein’s
quaternary structure.
In this study, we focus on protein homodimers — a common
quaternary structural motif. The Protein Data Bank (PDB)
was surveyed for protein dimers and for each individual
entry, monomeric dipoles were calculated and analyzed using
the program GRASP. A highly significant number of dimers
were characterized by monomeric dipole moments that were
oriented in opposite directions implying that the twofold
symmetry element that generates the dimer is
perpendicular to the dipole moment axis. This results in
a net dipole moment for the overall dimer that is close
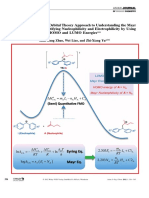
to zero. Two examples — transforming growth factor
β1 and the HIV protease enzyme — are depicted here
(the dipole from one monomer is shown in red with
the dipole from the corresponding symmetry mate in
blue). Quantitative analyses of dipole moments and
their symmetry relationships to the dimer twofold
also confirm these observations.
A number of conclusions flow from this result. The
answer as to why proteins oligomerize is multiple. As
is well known, oligomerization serves to make
cooperativity and allostery possible and also serves to
decrease the colloid properties (osmotic tension) of
cytoplasmic and extracellular protein concentrations.
To these major reasons we can also add a third —
proteins dimerize in order to minimize the overall, net molecular dipole and thus minimize the largescale
electrostatic energy. This result would suggest a general principle of quaternary structure. Furthermore,
protein oligomerization processes involved in signal transduction and transcriptional regulation may also be
determined by such intermolecular dipole interactions. Small conformational changes that modify the
dipole moment of one monomer may thus cause largescale reorientations of protein oligomers. In a fashion
similar to this work, further study of such higherorder protein structures may confirm this hypothesis.
Você também pode gostar
- Deciphering Hidden DNA Meta-Codes - The Great Unification & Master Code of BiologyDocumento20 páginasDeciphering Hidden DNA Meta-Codes - The Great Unification & Master Code of BiologyJean-claude PerezAinda não há avaliações
- Lehninger Ch26Documento81 páginasLehninger Ch26AMAN KUMAR SINGH100% (1)
- Biology Staar Review Stations Day 3Documento12 páginasBiology Staar Review Stations Day 3api-267841335Ainda não há avaliações
- Chem 40.1 PostlabDocumento6 páginasChem 40.1 PostlabaraneyaAinda não há avaliações
- Zinc Finger Proteins From Atomic Contact To Cellular FunctionDocumento292 páginasZinc Finger Proteins From Atomic Contact To Cellular FunctionGsureshbabuAinda não há avaliações
- Introducing Protein Folding Using Simple ModelsDocumento32 páginasIntroducing Protein Folding Using Simple Modelstestonly261Ainda não há avaliações
- Protein Dynamics and Enzyme Catalysis - Insights From SimulationsDocumento16 páginasProtein Dynamics and Enzyme Catalysis - Insights From Simulationsyin chanAinda não há avaliações
- Wecb 354Documento10 páginasWecb 354azzaassAinda não há avaliações
- Maisuradze 2012Documento16 páginasMaisuradze 2012Geysel SuarezAinda não há avaliações
- The Kinetics of Nucleated Polymerizations at High Concentrations: Amyloid Fibril Formation Near and Above The Supercritical Concentration''Documento11 páginasThe Kinetics of Nucleated Polymerizations at High Concentrations: Amyloid Fibril Formation Near and Above The Supercritical Concentration''Fatima Herranz TrilloAinda não há avaliações
- Genetic Algorithm For Protein Folding Simulations in 2D HP ModelDocumento11 páginasGenetic Algorithm For Protein Folding Simulations in 2D HP ModelToño GonzálezAinda não há avaliações
- Stochasticity in Gene Expression From Theories To PhenotypesDocumento14 páginasStochasticity in Gene Expression From Theories To PhenotypesNando93Ainda não há avaliações
- TMP 70 D3Documento20 páginasTMP 70 D3FrontiersAinda não há avaliações
- Amino Acid ClassificationDocumento15 páginasAmino Acid ClassificationRithushreeAinda não há avaliações
- Macromolecular CrowdingDocumento10 páginasMacromolecular CrowdingChris McMahonAinda não há avaliações
- Research Proposal Revised FinalDocumento5 páginasResearch Proposal Revised Finalapi-300595323Ainda não há avaliações
- De Jesus-Allen-Bba Biomembranes-2013 - Determinants-Of-Hyrophobic-MismatchDocumento13 páginasDe Jesus-Allen-Bba Biomembranes-2013 - Determinants-Of-Hyrophobic-MismatchLester John VeraAinda não há avaliações
- Hall 1979Documento6 páginasHall 1979sepot24093Ainda não há avaliações
- J. Biol. Chem.-1999-Xie-15967-70Documento4 páginasJ. Biol. Chem.-1999-Xie-15967-70ashueinAinda não há avaliações
- Statistical Measures To Quantify Similarity Between Molecular Dynamics Simulation TrajectoriesDocumento17 páginasStatistical Measures To Quantify Similarity Between Molecular Dynamics Simulation TrajectoriesDiego GranadosAinda não há avaliações
- Jmc105061 (Molecular Interactions)Documento24 páginasJmc105061 (Molecular Interactions)Placido A. Ceballos ChiarucciAinda não há avaliações
- Recent Applications of Kirkwood-Buff Theory To Biological SystemsDocumento22 páginasRecent Applications of Kirkwood-Buff Theory To Biological SystemsanshuAinda não há avaliações
- Antypov 2007Documento8 páginasAntypov 2007brouuorbAinda não há avaliações
- HardwareDocumento16 páginasHardwarepaulogameiroAinda não há avaliações
- Poing PDFDocumento16 páginasPoing PDFAanAinda não há avaliações
- 1 s2.0 S0969212611003364 MainDocumento11 páginas1 s2.0 S0969212611003364 Maintiago almeidaAinda não há avaliações
- Biomolecules: Quantum Mechanical Modeling: A Tool For The Understanding of Enzyme ReactionsDocumento41 páginasBiomolecules: Quantum Mechanical Modeling: A Tool For The Understanding of Enzyme ReactionsCarlos Alberto Dutra Fraga FilhoAinda não há avaliações
- J of Molecular Recognition - 1999 - Jelesarov - Isothermal Titration Calorimetry and Differential Scanning Calorimetry AsDocumento16 páginasJ of Molecular Recognition - 1999 - Jelesarov - Isothermal Titration Calorimetry and Differential Scanning Calorimetry AscaropsAinda não há avaliações
- Using Protein Chemistry As A Way Into Teaching Chemical Bonding To ADocumento30 páginasUsing Protein Chemistry As A Way Into Teaching Chemical Bonding To ABernadeth UrsuaAinda não há avaliações
- 41 Models For Enzyme ActionDocumento10 páginas41 Models For Enzyme Actiontri sutrianiAinda não há avaliações
- MainDocumento9 páginasMainpaxjr2Ainda não há avaliações
- Post Translational Modifications of Proteins - ReviewDocumento20 páginasPost Translational Modifications of Proteins - ReviewNishita GuptaAinda não há avaliações
- Computational Enzymology: Methods in Molecular Biology (Clifton, N.J.) January 2013Documento25 páginasComputational Enzymology: Methods in Molecular Biology (Clifton, N.J.) January 2013detki007Ainda não há avaliações
- P2 Phar ChemDocumento22 páginasP2 Phar ChemFitri AinunAinda não há avaliações
- Interactions of Cytochrome P450s With Their LigandsDocumento10 páginasInteractions of Cytochrome P450s With Their LigandshlpcravoAinda não há avaliações
- Molecular Dynamics Simulations of Membrane Proteins: Philip C. Biggin and Peter J. BondDocumento18 páginasMolecular Dynamics Simulations of Membrane Proteins: Philip C. Biggin and Peter J. BondKübra KahveciAinda não há avaliações
- Full PaperDocumento10 páginasFull PaperĐình Thư LêAinda não há avaliações
- 0039-Bh-Bio-T-19 (Muhammad Ahsan)Documento9 páginas0039-Bh-Bio-T-19 (Muhammad Ahsan)خاک نشینAinda não há avaliações
- Postraductional Prot Review 2021Documento20 páginasPostraductional Prot Review 2021Diana CortesAinda não há avaliações
- All-Atom Molecular Dynamics Studies of The Full-Length B - Amyloid PeptidesDocumento10 páginasAll-Atom Molecular Dynamics Studies of The Full-Length B - Amyloid PeptidesOliverAinda não há avaliações
- Docking Chapter ProofDocumento26 páginasDocking Chapter ProofbiommcellAinda não há avaliações
- Protein To D2O InducesDocumento20 páginasProtein To D2O InducesibrahimAinda não há avaliações
- Burgess 2014Documento25 páginasBurgess 2014Essy MutiaaraAinda não há avaliações
- Size and Shape of Proteins - KDa To NMDocumento20 páginasSize and Shape of Proteins - KDa To NMElena Rojo de BenitoAinda não há avaliações
- tmp3539 TMPDocumento10 páginastmp3539 TMPFrontiersAinda não há avaliações
- Biology 1002B Outcomes: Basic Characteristics of Life (Chapter 2)Documento6 páginasBiology 1002B Outcomes: Basic Characteristics of Life (Chapter 2)Jenny JooAinda não há avaliações
- 11.hemocompatibility Assessment of PDMAEMA - Cerda.jcrDocumento10 páginas11.hemocompatibility Assessment of PDMAEMA - Cerda.jcrRebeca FloresAinda não há avaliações
- RNA and Protein 3D Structure Modeling: Similarities and DifferencesDocumento12 páginasRNA and Protein 3D Structure Modeling: Similarities and DifferencescsAinda não há avaliações
- 1409 2406 PDFDocumento21 páginas1409 2406 PDFGargi SharmaAinda não há avaliações
- Notes From Someone OnlineDocumento50 páginasNotes From Someone OnlineweldeenytAinda não há avaliações
- A. 1992. Makhatadze, G. I. Protein Interactions With Urea and Guanidium ChlorideDocumento15 páginasA. 1992. Makhatadze, G. I. Protein Interactions With Urea and Guanidium ChlorideJorge Luis RomeroAinda não há avaliações
- Artigo Mostra Glicosilação No RetículoDocumento20 páginasArtigo Mostra Glicosilação No RetículoMsDebora ReinertAinda não há avaliações
- Mass Spectrometry and The Age of The ProteomeDocumento19 páginasMass Spectrometry and The Age of The ProteomeNeshat HaqAinda não há avaliações
- Biomolecules: Kinetics and Thermodynamics of Membrane Protein FoldingDocumento20 páginasBiomolecules: Kinetics and Thermodynamics of Membrane Protein Foldingdon boscoAinda não há avaliações
- (2023) (Thoudam) Effect of Topology On The Collapse Transition and The Instantaneous Shape of A Model HeteropolymerDocumento13 páginas(2023) (Thoudam) Effect of Topology On The Collapse Transition and The Instantaneous Shape of A Model HeteropolymerIsabel CamusAinda não há avaliações
- Gardiner 2001Documento13 páginasGardiner 2001taoufik akabliAinda não há avaliações
- 10-Molecular Recognition ModelsDocumento7 páginas10-Molecular Recognition ModelsAnamarijaAinda não há avaliações
- Chromatophore Manuscript Final 03 24 19Documento25 páginasChromatophore Manuscript Final 03 24 19SigmundAinda não há avaliações
- 2064 Lingin 3layerDocumento13 páginas2064 Lingin 3layerM IdreesAinda não há avaliações
- Research Papers On Protein FoldingDocumento5 páginasResearch Papers On Protein Foldingshvximwhf100% (1)
- Generic CG Folding Aggre ModelsDocumento16 páginasGeneric CG Folding Aggre Modelssoumava palitAinda não há avaliações
- Learning Physical Properties of Organic Compounds Using Molecular ModelingDocumento9 páginasLearning Physical Properties of Organic Compounds Using Molecular ModelingUperma Kopma UnhasAinda não há avaliações
- Molecular Mechanisms of Direct and Indirect Interplay Between A - 2022 - ChemPhyDocumento11 páginasMolecular Mechanisms of Direct and Indirect Interplay Between A - 2022 - ChemPhyKetua PpkAinda não há avaliações
- RPD System IDocumento62 páginasRPD System IogangurelAinda não há avaliações
- MR 22 - Liberty and Equality - A Two-Part InventionDocumento9 páginasMR 22 - Liberty and Equality - A Two-Part InventionogangurelAinda não há avaliações
- Information Theoretic Correlates of Coding Storing and Learning in Neuronal SystemsDocumento37 páginasInformation Theoretic Correlates of Coding Storing and Learning in Neuronal SystemsogangurelAinda não há avaliações
- MR 22 - Lecture Notes Fall 1983Documento97 páginasMR 22 - Lecture Notes Fall 1983ogangurelAinda não há avaliações
- MR 22 - Contracts and The Murder of Roger WhetmoreDocumento8 páginasMR 22 - Contracts and The Murder of Roger WhetmoreogangurelAinda não há avaliações
- RPD System 2Documento35 páginasRPD System 2ogangurelAinda não há avaliações
- Scientific Conceptualization of InformationDocumento24 páginasScientific Conceptualization of InformationogangurelAinda não há avaliações
- Extraction Experiments On Coated VesiclesDocumento19 páginasExtraction Experiments On Coated VesiclesogangurelAinda não há avaliações
- CS91r Jan 84 Integrated Support Environments For Hierarchical Software DevelopmentDocumento41 páginasCS91r Jan 84 Integrated Support Environments For Hierarchical Software DevelopmentogangurelAinda não há avaliações
- Congressional Record EntryDocumento1 páginaCongressional Record EntryogangurelAinda não há avaliações
- CS91r May 84 Meta-Language Based Environments For Integrated Software Development PT 2Documento12 páginasCS91r May 84 Meta-Language Based Environments For Integrated Software Development PT 2ogangurelAinda não há avaliações
- APLSIMULDocumento39 páginasAPLSIMULogangurelAinda não há avaliações
- CS91r May 84 Meta-Language Based Environments For Integrated Software Development PT 1Documento44 páginasCS91r May 84 Meta-Language Based Environments For Integrated Software Development PT 1ogangurelAinda não há avaliações
- CONGRESSIONAL RECORD - Extensions of Remarks E29: January 6, 2011Documento3 páginasCONGRESSIONAL RECORD - Extensions of Remarks E29: January 6, 2011ogangurelAinda não há avaliações
- Complete Stories - Word VersionDocumento92 páginasComplete Stories - Word VersionogangurelAinda não há avaliações
- WV - PennsylvaniaDocumento4 páginasWV - PennsylvaniaogangurelAinda não há avaliações
- Kucinich CitationDocumento2 páginasKucinich CitationogangurelAinda não há avaliações
- Illinois - IndianaDocumento3 páginasIllinois - IndianaogangurelAinda não há avaliações
- Hagerstown Herald-MailDocumento3 páginasHagerstown Herald-MailogangurelAinda não há avaliações
- Maryland - DCDocumento2 páginasMaryland - DCogangurelAinda não há avaliações
- The Glow of Life - Whole Body ScanningDocumento2 páginasThe Glow of Life - Whole Body Scanningogangurel100% (1)
- Media Coverage For The Walk For HealthcareDocumento8 páginasMedia Coverage For The Walk For HealthcareogangurelAinda não há avaliações
- Table of ContentsDocumento1 páginaTable of Contentsogangurel100% (2)
- Lima NewsDocumento2 páginasLima NewsogangurelAinda não há avaliações
- Walk For Healthcare: Stories From MarylandDocumento7 páginasWalk For Healthcare: Stories From MarylandogangurelAinda não há avaliações
- Walk For Healthcare: Stories From PennsylvaniaDocumento14 páginasWalk For Healthcare: Stories From PennsylvaniaogangurelAinda não há avaliações
- Walk For Healthcare: Stories From OhioDocumento16 páginasWalk For Healthcare: Stories From OhioogangurelAinda não há avaliações
- Real People With Real Stories From The Walk For HealthcareDocumento61 páginasReal People With Real Stories From The Walk For HealthcareogangurelAinda não há avaliações
- Walk For Healthcare: Stories From IndianaDocumento14 páginasWalk For Healthcare: Stories From IndianaogangurelAinda não há avaliações
- "Share Your Story" Event at Chen's in Chicago, Thursday 8-27Documento1 página"Share Your Story" Event at Chen's in Chicago, Thursday 8-27ogangurelAinda não há avaliações
- Written Exam in Molecular Modeling: InstructionsDocumento11 páginasWritten Exam in Molecular Modeling: InstructionsTalal Ahmed Awad MohammedAinda não há avaliações
- DNA Replication 2014Documento15 páginasDNA Replication 2014SATYANARAYANA RAO100% (1)
- Stryer Table of ContentsDocumento5 páginasStryer Table of ContentsHannah Grace A PugalAinda não há avaliações
- Dna ReplicationDocumento46 páginasDna ReplicationThomas AbichAinda não há avaliações
- MMC 3Documento27 páginasMMC 3oli.finetAinda não há avaliações
- Chapter 22 BiochemistryDocumento13 páginasChapter 22 Biochemistrynamini20Ainda não há avaliações
- Gene SilencingDocumento7 páginasGene SilencingMaliha JahanAinda não há avaliações
- Gene RegulationDocumento15 páginasGene RegulationRajendra PrajapatiAinda não há avaliações
- SY 2022-2023 Updated Chem 301 Biochem Lec Synch and AsynchDocumento3 páginasSY 2022-2023 Updated Chem 301 Biochem Lec Synch and AsynchLYKA ANTONETTE ABREGANAAinda não há avaliações
- Science of SlimeDocumento3 páginasScience of SlimeMax Is hereAinda não há avaliações
- Enzyme Promiscuity in Earthworm Serine Protease: Substrate Versatility and Therapeutic PotentialDocumento8 páginasEnzyme Promiscuity in Earthworm Serine Protease: Substrate Versatility and Therapeutic PotentialKhánh HuyềnAinda não há avaliações
- St. Thomas Academy: Pob. 3, City of Sto. Tomas, Batangas SY 2021 - 2022Documento4 páginasSt. Thomas Academy: Pob. 3, City of Sto. Tomas, Batangas SY 2021 - 2022Kate Ashly Mae AteradoAinda não há avaliações
- Vitis OutputDocumento3.573 páginasVitis OutputKian Bigovic VilliAinda não há avaliações
- Bio 024 - Quiz Cfu Sas 3 (Answer Key)Documento4 páginasBio 024 - Quiz Cfu Sas 3 (Answer Key)ELLE WOODSAinda não há avaliações
- RIBOZYMESDocumento17 páginasRIBOZYMESSharan GayathrinathanAinda não há avaliações
- Lecture Notes - Weeks 1 and 2 BCMB2001Documento17 páginasLecture Notes - Weeks 1 and 2 BCMB2001Anushka NeupaneAinda não há avaliações
- MCB RevisionDocumento73 páginasMCB RevisionRitika SahniAinda não há avaliações
- P GEXmapDocumento1 páginaP GEXmapAnonymous byoeVOAinda não há avaliações
- Heredity Dna RnaDocumento45 páginasHeredity Dna RnariddhiAinda não há avaliações
- CBSE Class 11 Biology Biomolecules NotesDocumento4 páginasCBSE Class 11 Biology Biomolecules NoteslokpatiAinda não há avaliações
- Biology Note For Grade 12 (Unit Three Molecular GeneticsDocumento24 páginasBiology Note For Grade 12 (Unit Three Molecular GeneticsDaniel GtsadkanAinda não há avaliações
- Restriction EnzymeDocumento33 páginasRestriction EnzymeFawaz MohamadAinda não há avaliações
- Mass Production of Bovine Serum AlbuminDocumento4 páginasMass Production of Bovine Serum AlbuminInternational Journal of Innovative Science and Research TechnologyAinda não há avaliações
- Chemistry Notes For Class 12 Chapter 14 Biomolecules: CarbohydratesDocumento20 páginasChemistry Notes For Class 12 Chapter 14 Biomolecules: CarbohydratesSOUMYODEEP NAYAKAinda não há avaliações
- Test Nucleic Acids SLDocumento2 páginasTest Nucleic Acids SLKatrīna SimanovskaAinda não há avaliações
- PROTEIN (Polypeptide) : Protein: Senyawa Organik Yang Merupakan Polimer Asam Amino Penyusun ProteinDocumento79 páginasPROTEIN (Polypeptide) : Protein: Senyawa Organik Yang Merupakan Polimer Asam Amino Penyusun ProteinDwinur ChasanahAinda não há avaliações